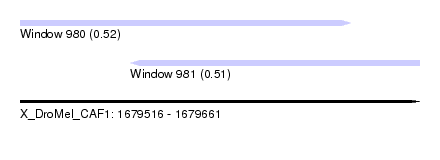
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,679,516 – 1,679,661 |
| Length | 145 |
| Max. P | 0.516338 |

| Location | 1,679,516 – 1,679,636 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.59 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -22.22 |
| Energy contribution | -21.67 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.79 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516338 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
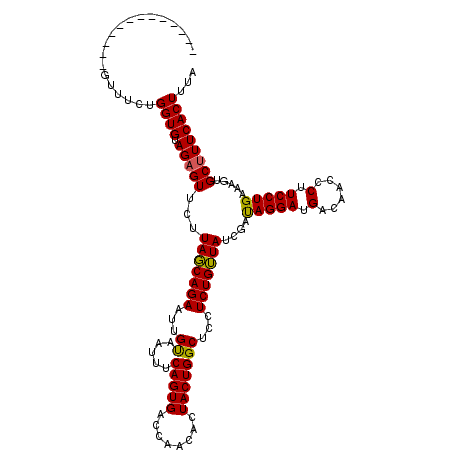
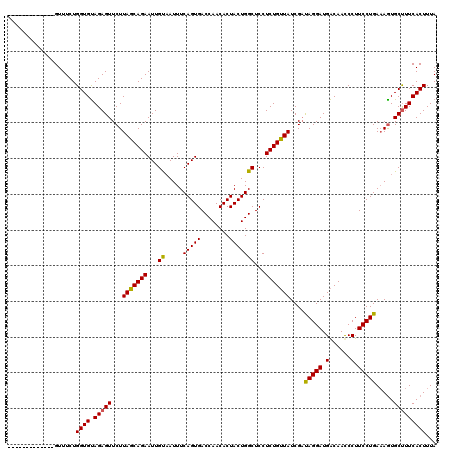
>X_DroMel_CAF1 1679516 120 + 22224390 CUAUCACGUUAACGUUUCUGGUGCAGAGUUCUUACCAGAAUUGCCAUUUCAGUGACCAACACUACUGGCUCCUCUGGUAUCGUCAGGAUGGCAACCCUUCCUGAUAGUGCCUUCACUUUA .....(((....)))((((((((.((....))))))))))..((((....((((.....))))..))))......(((((..((((((.((....)).))))))..)))))......... ( -37.80) >DroSec_CAF1 19747 107 + 1 -------------GUUUCUGGUGUAGAGUUCUUAGCAGAAUUGUAAUUUCAGUGACCAACACUACUGGCUCCUCUGUUAUCGAUAGGAUGACAAUCCUUCCUGAAAGUGCUUUCACUUUA -------------......((((.(((((.(((((((((...((......((((.....))))....))...)))))).(((..(((((....)))))...)))))).)))))))))... ( -23.90) >DroSim_CAF1 13599 107 + 1 -------------GUUUCUGGUGUAGAGUUCUUAGCAGAAUUGUAAUUUCAGUGACCAACACUACUGGCUCCUCUGUUAUCGAUAGGAUGACAACCCUUCCUGAAAAUGCUUUCACUUUA -------------..(((((.((.((....)))).)))))..(((.((((((.(((((.......)))......(((((((.....))))))).....)))))))).))).......... ( -22.20) >consensus _____________GUUUCUGGUGUAGAGUUCUUAGCAGAAUUGUAAUUUCAGUGACCAACACUACUGGCUCCUCUGUUAUCGAUAGGAUGACAACCCUUCCUGAAAGUGCUUUCACUUUA ...................((((.(((((...(((((((...((.....(((((........)))))))...)))))))....(((((.(......).))))).....)))))))))... (-22.22 = -21.67 + -0.55)

| Location | 1,679,556 – 1,679,661 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.44 |
| Mean single sequence MFE | -29.73 |
| Consensus MFE | -19.59 |
| Energy contribution | -19.27 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.505359 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
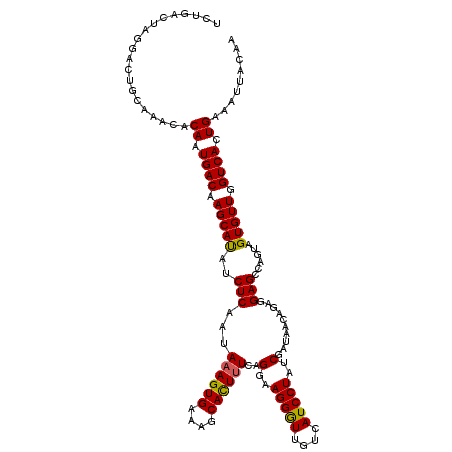
>X_DroMel_CAF1 1679556 105 - 22224390 ---------------AAACACAAUGACGAGCACUUCUCAAUAAAGUGAAGGCACUAUCAGGAAGGGUUGCCAUCCUGACGAUACCAGAGGAGCCAGUAGUGUUGGUCACUGAAAUGGCAA ---------------.....((.((((.((((((.((((......))).(((....((((((.((....)).)))))).....((...)).))).).)))))).)))).))......... ( -30.40) >DroSec_CAF1 19774 120 - 1 UCUGACUAGGACUGCAAACACAAUGACAAGCAUAUCUCAAUAAAGUGAAAGCACUUUCAGGAAGGAUUGUCAUCCUAUCGAUAACAGAGGAGCCAGUAGUGUUGGUCACUGAAAUUACAA ..(((((((.(((((........((((((.(...(((....((((((....))))))..)))..).))))))((((...........))))....))))).)))))))............ ( -29.50) >DroSim_CAF1 13626 120 - 1 UCUGACUAGGACUGCAAACACAAUGACAAGCAUAUCUCAAUAAAGUGAAAGCAUUUUCAGGAAGGGUUGUCAUCCUAUCGAUAACAGAGGAGCCAGUAGUGUUGGUCACUGAAAUUACAA ..(((((((.(((((........((((((.(...(((....((((((....))))))..)))..).))))))((((...........))))....))))).)))))))............ ( -29.30) >consensus UCUGACUAGGACUGCAAACACAAUGACAAGCAUAUCUCAAUAAAGUGAAAGCACUUUCAGGAAGGGUUGUCAUCCUAUCGAUAACAGAGGAGCCAGUAGUGUUGGUCACUGAAAUUACAA ....................((.((((.(((((..(((...((((((....))))))..(..(((((....)))))..)..........)))......))))).)))).))......... (-19.59 = -19.27 + -0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:49:16 2006