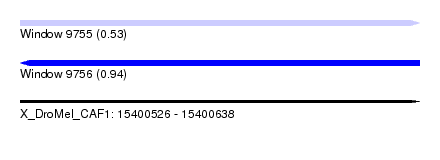

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,400,526 – 15,400,638 |
| Length | 112 |
| Max. P | 0.936890 |

| Location | 15,400,526 – 15,400,638 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 70.14 |
| Mean single sequence MFE | -28.75 |
| Consensus MFE | -11.85 |
| Energy contribution | -11.85 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.41 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532323 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15400526 112 + 22224390 AUUUUAUGUAAUACU-----GUGGCGCGGCAUA-UUUUGUUGUGAACAAUGAAAUGCGUUAAUUCGAGGGG-UUUGGUUCAGCGAAAAGCCAAUUUAAUUGUGGCAUUCGCCACACUUG ...............-----.((((..(((.((-(((..(((....)))..))))).)))..((((..(..-(...)..)..))))..)))).......((((((....)))))).... ( -29.30) >DroSec_CAF1 46186 103 + 1 ---------AAAAUU-----GUGGCGCGGCAUA-UUUUGUUGUGAACAAUAAAAUGCGUUAAUUCGAGGGG-UUUGCUUCAGCGAAAAGCCAAUUUAAUUGUGGCAUUCGCCACACUUG ---------.....(-----((((((((((.((-((((((((....)))))))))).))).....((((..-....)))).))(((..((((.........)))).))))))))).... ( -31.70) >DroEre_CAF1 37843 93 + 1 AAC---------UCC-----GUGGCGCGGCAUA-UUUGCU------CAA-AAAAUGCCUUAAUUUGAGGGG-UUUGGCUCAGCGAAAAGCCAAUUUCAUUGUGGUAUUCAGCACAC--- ...---------...-----(((((..((((..-..))))------...-..((((((.((((....(..(-((.((((........)))))))..))))).))))))..)).)))--- ( -21.80) >DroYak_CAF1 41028 101 + 1 AAU---------UCUAUUUUGUUGUGAGCAAUAAACUGCU------CAUUAAAAUGCGUUAAUUUGAUGGUUUUUGGCUCAGCGAAGAGCCAAUUUCAUUGUGGUAUUUAACGCAU--- ...---------...((((((..(((((((......))))------)))))))))(((((((..((((((...(((((((......)))))))..))))))......)))))))..--- ( -32.20) >consensus AAU_________ACU_____GUGGCGCGGCAUA_UUUGCU______CAAUAAAAUGCGUUAAUUCGAGGGG_UUUGGCUCAGCGAAAAGCCAAUUUAAUUGUGGCAUUCACCACAC___ ....................((((((....................................((((..(((......)))..))))..((((.........))))...))))))..... (-11.85 = -11.85 + -0.00)


| Location | 15,400,526 – 15,400,638 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 70.14 |
| Mean single sequence MFE | -23.33 |
| Consensus MFE | -10.95 |
| Energy contribution | -11.33 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.39 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.47 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.936890 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
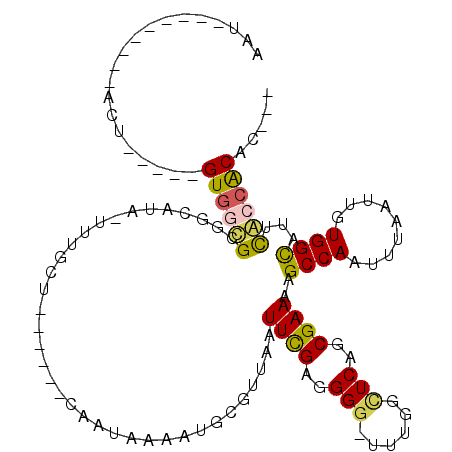
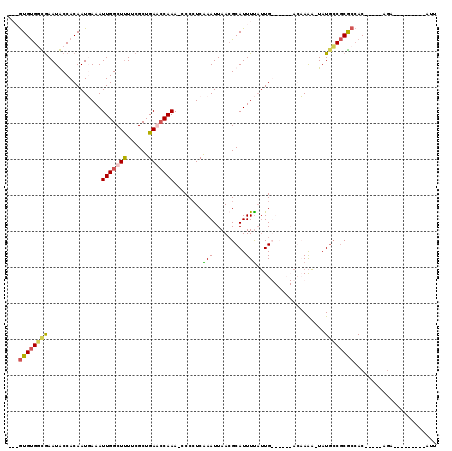
>X_DroMel_CAF1 15400526 112 - 22224390 CAAGUGUGGCGAAUGCCACAAUUAAAUUGGCUUUUCGCUGAACCAAA-CCCCUCGAAUUAACGCAUUUCAUUGUUCACAACAAAA-UAUGCCGCGCCAC-----AGUAUUACAUAAAAU ..(.(((((((((.((((.........))))..)))(((((......-....))).......((((....((((.....))))..-.)))).)))))))-----).)............ ( -22.00) >DroSec_CAF1 46186 103 - 1 CAAGUGUGGCGAAUGCCACAAUUAAAUUGGCUUUUCGCUGAAGCAAA-CCCCUCGAAUUAACGCAUUUUAUUGUUCACAACAAAA-UAUGCCGCGCCAC-----AAUUUU--------- ...(((((((((((((..((((...))))(((((.....)))))...-..............))))))..((((.....))))..-...)))))))...-----......--------- ( -22.60) >DroEre_CAF1 37843 93 - 1 ---GUGUGCUGAAUACCACAAUGAAAUUGGCUUUUCGCUGAGCCAAA-CCCCUCAAAUUAAGGCAUUUU-UUG------AGCAAA-UAUGCCGCGCCAC-----GGA---------GUU ---((((((.................(((((((......))))))).-.............(((((.((-(..------...)))-.)))))))).)))-----...---------... ( -22.10) >DroYak_CAF1 41028 101 - 1 ---AUGCGUUAAAUACCACAAUGAAAUUGGCUCUUCGCUGAGCCAAAAACCAUCAAAUUAACGCAUUUUAAUG------AGCAGUUUAUUGCUCACAACAAAAUAGA---------AUU ---(((((((((....(.....)...(((((((......)))))))...........))))))))).....((------(((((....)))))))............---------... ( -26.60) >consensus ___GUGUGGCGAAUACCACAAUGAAAUUGGCUUUUCGCUGAACCAAA_CCCCUCAAAUUAACGCAUUUUAUUG______ACAAAA_UAUGCCGCGCCAC_____AGA_________AUU ...((((((((...............(((((((......)))))))........(((.........)))...................))))))))....................... (-10.95 = -11.33 + 0.38)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:47 2006