
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,238,280 – 15,238,393 |
| Length | 113 |
| Max. P | 0.922002 |

| Location | 15,238,280 – 15,238,393 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 82.39 |
| Mean single sequence MFE | -27.91 |
| Consensus MFE | -22.07 |
| Energy contribution | -22.23 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.712884 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15238280 113 + 22224390 GUGCGUACAUAUAU-CUGCGGAUGACCCCAUUGUUGUUU-UUGGGGUCAGCUG-AACCAUUUGUGGACUUUCGAUUUAAUAUAAAUUGAGAAUGAGCUUGUUAGUUUUGGCGCUUG ((((((.((((...-...(((.((((((((..(....).-.)))))))).)))-.......)))).))((((((((((...))))))))))..(((((....)))))..))))... ( -32.82) >DroPse_CAF1 64687 88 + 1 GUGCGU-----------GUGGGUGACCCCAUUGUUGUU--UUGGGGUCAGCUG-AACCAUUUGUGGACUUUCGAUUUAAUAUAAAUUGGGAAUGAGCUU--------------UUC ..((((-----------..((.((((((((........--.)))))))).)).-..(((....)))))(..(((((((...)))))))..)....))..--------------... ( -25.10) >DroSec_CAF1 58600 109 + 1 GUGCGU----AUAU-CUGUGGAUGACCCCAUUGUUGUUU-UUGGGGUCAGCUG-AACCAUUUGUGAACUUUCGAUUUAAUAUAAAUUGAGAAUGAGCUUGUUAGUUUUGGCGCUUG ((((((----.(((-..((((.((((((((..(....).-.))))))))....-..))))..))).))((((((((((...))))))))))..(((((....)))))..))))... ( -30.80) >DroYak_CAF1 59364 109 + 1 GUGCGU----AUAU-CUGCGGAUGACCCCAUUGUUGUUU-UUGGGGUCAGCUG-AACCAUUUGUGGACUUUCGAUUUAAUAUAAAUUGCGAAUGAGCUUGUUAGUUUUGGCGCUUG ..((((----....-((((((.((((((((..(....).-.)))))))).)))-...((((((..(...................)..)))))).......))).....))))... ( -28.21) >DroAna_CAF1 59484 100 + 1 -UGCGU----GUAUGCUG----UGACCCCAUUGUUGUUUUUUGGGGUCAGCUGGAGCCAUUUGUGGACUUUCGAUUUAAUAUAAAUUGAGAAUGAGCUUGUUAGUUUUG------- -.(((.----...)))..----((((((((..(.....)..))))))))(((((((((((((........((((((((...))))))))))))).))))..))))....------- ( -25.40) >DroPer_CAF1 66400 88 + 1 GUGCGU-----------GUGGGUGACCCCAUUGUUGUU--UUGGGGUCAGCUG-AACCAUUUGUGGACUUUCGAUUUAAUAUAAAUUGGGAAUGAGCUU--------------UUC ..((((-----------..((.((((((((........--.)))))))).)).-..(((....)))))(..(((((((...)))))))..)....))..--------------... ( -25.10) >consensus GUGCGU____AUAU_CUGUGGAUGACCCCAUUGUUGUUU_UUGGGGUCAGCUG_AACCAUUUGUGGACUUUCGAUUUAAUAUAAAUUGAGAAUGAGCUUGUUAGUUUUG____UUG ......................((((((((...........))))))))(((....(((....)))..((((((((((...))))))))))...)))................... (-22.07 = -22.23 + 0.17)

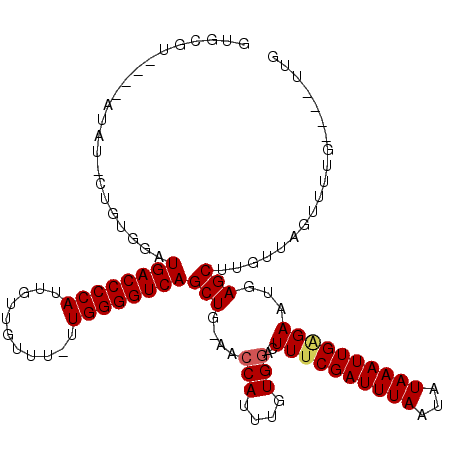

| Location | 15,238,280 – 15,238,393 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 82.39 |
| Mean single sequence MFE | -18.88 |
| Consensus MFE | -16.04 |
| Energy contribution | -15.90 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.922002 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15238280 113 - 22224390 CAAGCGCCAAAACUAACAAGCUCAUUCUCAAUUUAUAUUAAAUCGAAAGUCCACAAAUGGUU-CAGCUGACCCCAA-AAACAACAAUGGGGUCAUCCGCAG-AUAUAUGUACGCAC ...(((............((((.....................(....).(((....)))..-.))))(((((((.-.........)))))))....(((.-.....))).))).. ( -20.80) >DroPse_CAF1 64687 88 - 1 GAA--------------AAGCUCAUUCCCAAUUUAUAUUAAAUCGAAAGUCCACAAAUGGUU-CAGCUGACCCCAA--AACAACAAUGGGGUCACCCAC-----------ACGCAC ...--------------.((((.....................(....).(((....)))..-.))))(((((((.--........)))))))......-----------...... ( -16.50) >DroSec_CAF1 58600 109 - 1 CAAGCGCCAAAACUAACAAGCUCAUUCUCAAUUUAUAUUAAAUCGAAAGUUCACAAAUGGUU-CAGCUGACCCCAA-AAACAACAAUGGGGUCAUCCACAG-AUAU----ACGCAC ...((((...(((((........(((.((.(((((...))))).)).))).......)))))-..))((((((((.-.........)))))))).......-....----..)).. ( -18.26) >DroYak_CAF1 59364 109 - 1 CAAGCGCCAAAACUAACAAGCUCAUUCGCAAUUUAUAUUAAAUCGAAAGUCCACAAAUGGUU-CAGCUGACCCCAA-AAACAACAAUGGGGUCAUCCGCAG-AUAU----ACGCAC ...(((.............((......))....((((((.....(((...(((....)))))-).((((((((((.-.........))))))))...)).)-))))----)))).. ( -23.10) >DroAna_CAF1 59484 100 - 1 -------CAAAACUAACAAGCUCAUUCUCAAUUUAUAUUAAAUCGAAAGUCCACAAAUGGCUCCAGCUGACCCCAAAAAACAACAAUGGGGUCA----CAGCAUAC----ACGCA- -------............((..........................((.(((....)))))...((((((((((...........))))))).----.)))....----..)).- ( -18.10) >DroPer_CAF1 66400 88 - 1 GAA--------------AAGCUCAUUCCCAAUUUAUAUUAAAUCGAAAGUCCACAAAUGGUU-CAGCUGACCCCAA--AACAACAAUGGGGUCACCCAC-----------ACGCAC ...--------------.((((.....................(....).(((....)))..-.))))(((((((.--........)))))))......-----------...... ( -16.50) >consensus CAA____CAAAACUAACAAGCUCAUUCUCAAUUUAUAUUAAAUCGAAAGUCCACAAAUGGUU_CAGCUGACCCCAA_AAACAACAAUGGGGUCAUCCACAG_AUAU____ACGCAC ..................((((.....................(....).(((....)))....))))(((((((...........)))))))....................... (-16.04 = -15.90 + -0.14)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:59:27 2006