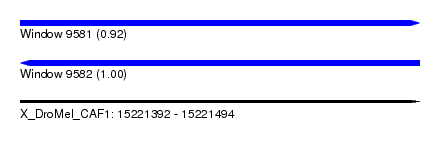
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,221,392 – 15,221,494 |
| Length | 102 |
| Max. P | 0.998551 |

| Location | 15,221,392 – 15,221,494 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 78.59 |
| Mean single sequence MFE | -20.12 |
| Consensus MFE | -12.95 |
| Energy contribution | -15.20 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.917505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15221392 102 + 22224390 UUAUUAAAAUGUACAUCAAUUUGUAUGUGUUCUUACCACGAAAGGUGCGAACACUCAUUCAAGAUAUUCAAACAGAUAUAGUAUCUAAUAUUUAAGUUUGAC ..........(((((......)))))((((((.((((......)))).)))))).............((((((((((((........))))))..)))))). ( -21.30) >DroSim_CAF1 41588 102 + 1 UUGUUAAAAUGUACAUAAAUUUGUAUGCGUUCGUACCACGAAAGGUGAGAACACUCAUUCAAGAUACUCAAAUAGAUAUAGUAUCUAAUAUUUAAGUUUGAC ..((((((..(((((......)))))..((((.((((......)))).)))).........(((((((...........)))))))..........)))))) ( -19.00) >DroEre_CAF1 43295 101 + 1 UCGUUAAAAUUUUCAUAAAUUGUAAUGCGUUCCUACCAUUUAAGGUAGAAAC-CUUUUUCAAAAUACUUGAAUUGAUAUAGUACCUAAUCCUAAAGGUUGAU .(((((.(((((....))))).))))).....(((((......))))).(((-(((((((((.....)))))..(((.(((...))))))..)))))))... ( -15.80) >DroYak_CAF1 41751 101 + 1 UCGUUAAAAUUUGCAUAAAUUUAAAUCUGUUCCUACCACAGAAGGUAGGAACACUUUUUCAAGAUACUUGAAUAGAUGUAAUAUCUAAUACUAA-CUUUGAC ..((((....((((((...........((((((((((......))))))))))((..((((((...)))))).))))))))....)))).....-....... ( -24.40) >consensus UCGUUAAAAUGUACAUAAAUUUGAAUGCGUUCCUACCACGAAAGGUAGGAACACUCAUUCAAGAUACUCAAAUAGAUAUAGUAUCUAAUACUAAAGUUUGAC ..((((((..................(((((((((((......))))))))))).......(((((((...........)))))))..........)))))) (-12.95 = -15.20 + 2.25)



| Location | 15,221,392 – 15,221,494 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 78.59 |
| Mean single sequence MFE | -23.57 |
| Consensus MFE | -15.69 |
| Energy contribution | -16.62 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.92 |
| Structure conservation index | 0.67 |
| SVM decision value | 3.14 |
| SVM RNA-class probability | 0.998551 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15221392 102 - 22224390 GUCAAACUUAAAUAUUAGAUACUAUAUCUGUUUGAAUAUCUUGAAUGAGUGUUCGCACCUUUCGUGGUAAGAACACAUACAAAUUGAUGUACAUUUUAAUAA ............(((((((...((((((.(((((....((......))((((((..(((......)))..))))))...))))).))))))...))))))). ( -25.20) >DroSim_CAF1 41588 102 - 1 GUCAAACUUAAAUAUUAGAUACUAUAUCUAUUUGAGUAUCUUGAAUGAGUGUUCUCACCUUUCGUGGUACGAACGCAUACAAAUUUAUGUACAUUUUAACAA .....(((((((...((((((...))))))))))))).....(((((.((((((..(((......)))..)))))).((((......)))))))))...... ( -21.00) >DroEre_CAF1 43295 101 - 1 AUCAACCUUUAGGAUUAGGUACUAUAUCAAUUCAAGUAUUUUGAAAAAG-GUUUCUACCUUAAAUGGUAGGAACGCAUUACAAUUUAUGAAAAUUUUAACGA ....((((........))))......(((.((((((...))))))....-(((((((((......))))))))).............)))............ ( -18.80) >DroYak_CAF1 41751 101 - 1 GUCAAAG-UUAGUAUUAGAUAUUACAUCUAUUCAAGUAUCUUGAAAAAGUGUUCCUACCUUCUGUGGUAGGAACAGAUUUAAAUUUAUGCAAAUUUUAACGA ((.((((-((.((((.((((.(((.((((.((((((...)))))).....(((((((((......))))))))))))).))))))))))).)))))).)).. ( -29.30) >consensus GUCAAACUUAAAUAUUAGAUACUAUAUCUAUUCAAGUAUCUUGAAAAAGUGUUCCCACCUUUCGUGGUAGGAACACAUACAAAUUUAUGUAAAUUUUAACAA ................(((((((...........))))))).......(((((((((((......))))))))))).......................... (-15.69 = -16.62 + 0.94)

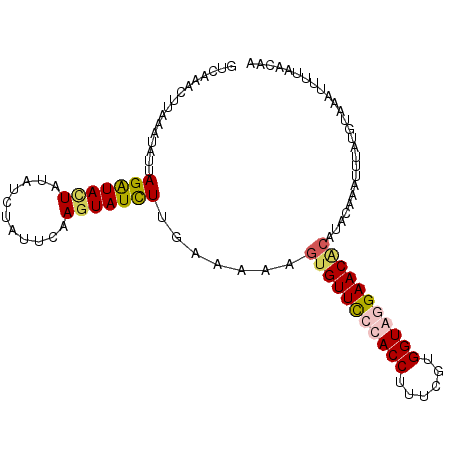

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:59:13 2006