| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,134,310 – 15,134,426 |
| Length | 116 |
| Max. P | 0.834730 |

| Location | 15,134,310 – 15,134,426 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 89.30 |
| Mean single sequence MFE | -34.90 |
| Consensus MFE | -25.01 |
| Energy contribution | -24.83 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834730 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15134310 116 + 22224390 ACAAAACAAGCGUGAAGAACUCG-AG--UCUCUCGACUUCGGCAUACCCAAAAAGCACGAAAGCGGGAAGUGUGAUUAUGGCCAAAAUUCCAGAUCUGCGCUUUAAUGGGUGUGCGAAG ............(((((...(((-((--...))))))))))(((((((((.....(.(....).).(((((((((((.(((........))))))).)))))))..))))))))).... ( -36.00) >DroSec_CAF1 45745 119 + 1 ACAAAACAAGCGAGAAGAACUCGGAGUCUCUCUCGACUUCGGCAUACCCAAAAAGCACGAAAGCGGGAAGUGUGAACAUGGCCAAAAUUCCAGAUCUGCGCUUUAAUGGGUGUGCGGAG ........((((((((((........))).))))).))((.(((((((((.(((((.(....)..((..(((....)))..))................)))))..))))))))).)). ( -38.20) >DroEre_CAF1 52668 110 + 1 --AAAACAAACC---AGAACUCG-AG--UCUAUCGACUUCGGUAUACCCAAAGACCACGAAAGCGGGAAAUGUGAAUAUGGCCAA-AUUCCAGAUCUGUGCUUUAAUGGGUGUGCGGAG --........((---.(((.(((-(.--....)))).))).(((((((((....((.(....).))((..((.((((........-))))))..))..........))))))))))).. ( -31.10) >DroYak_CAF1 53400 112 + 1 ACAAAACAAACC---AGAACUCG-AG--UCUAUCGACUGCGGCAUACCCAAAAAGCACGAAAGCGGAAAAUGUGAAUAUGGCCAA-AUUCCAGAUCUGCGCUUUAAUGGGUGUGCGGAG ..........((---......((-((--((....)))).))(((((((((.(((((......(((((...((.((((........-))))))..))))))))))..))))))))))).. ( -34.30) >consensus ACAAAACAAACC___AGAACUCG_AG__UCUAUCGACUUCGGCAUACCCAAAAAGCACGAAAGCGGGAAAUGUGAAUAUGGCCAA_AUUCCAGAUCUGCGCUUUAAUGGGUGUGCGGAG ....................................(((((.((((((((.(((((.(....).((((....((........))...))))........)))))..))))))))))))) (-25.01 = -24.83 + -0.19)



| Location | 15,134,310 – 15,134,426 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 89.30 |
| Mean single sequence MFE | -32.65 |
| Consensus MFE | -24.05 |
| Energy contribution | -24.67 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.622595 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15134310 116 - 22224390 CUUCGCACACCCAUUAAAGCGCAGAUCUGGAAUUUUGGCCAUAAUCACACUUCCCGCUUUCGUGCUUUUUGGGUAUGCCGAAGUCGAGAGA--CU-CGAGUUCUUCACGCUUGUUUUGU ....(((.(((((..((((((((((.(.((((.(.(((......))).).)))).).))).))))))).))))).))).(((.(((((...--))-))).)))................ ( -33.30) >DroSec_CAF1 45745 119 - 1 CUCCGCACACCCAUUAAAGCGCAGAUCUGGAAUUUUGGCCAUGUUCACACUUCCCGCUUUCGUGCUUUUUGGGUAUGCCGAAGUCGAGAGAGACUCCGAGUUCUUCUCGCUUGUUUUGU ....(((.(((((..((((((((((.(.((((.(.(((......))).).)))).).))).))))))).))))).)))..(((.((((((..((.....))..)))))))))....... ( -36.20) >DroEre_CAF1 52668 110 - 1 CUCCGCACACCCAUUAAAGCACAGAUCUGGAAU-UUGGCCAUAUUCACAUUUCCCGCUUUCGUGGUCUUUGGGUAUACCGAAGUCGAUAGA--CU-CGAGUUCU---GGUUUGUUUU-- ....((............))(((((((.(((((-((((...((((.((.....((((....))))..(((((.....))))))).))))..--.)-))))))))---)))))))...-- ( -25.60) >DroYak_CAF1 53400 112 - 1 CUCCGCACACCCAUUAAAGCGCAGAUCUGGAAU-UUGGCCAUAUUCACAUUUUCCGCUUUCGUGCUUUUUGGGUAUGCCGCAGUCGAUAGA--CU-CGAGUUCU---GGUUUGUUUUGU ..(((((.(((((..(((((((..(..(((((.-.((..........))..)))))..)..))))))).))))).)))((.((((....))--))-))......---)).......... ( -35.50) >consensus CUCCGCACACCCAUUAAAGCGCAGAUCUGGAAU_UUGGCCAUAUUCACACUUCCCGCUUUCGUGCUUUUUGGGUAUGCCGAAGUCGAGAGA__CU_CGAGUUCU___CGCUUGUUUUGU ....(((.(((((..((((((((((.(.((((...(((......)))...)))).).))).))))))).))))).))).(((.(((..........))).)))................ (-24.05 = -24.67 + 0.63)

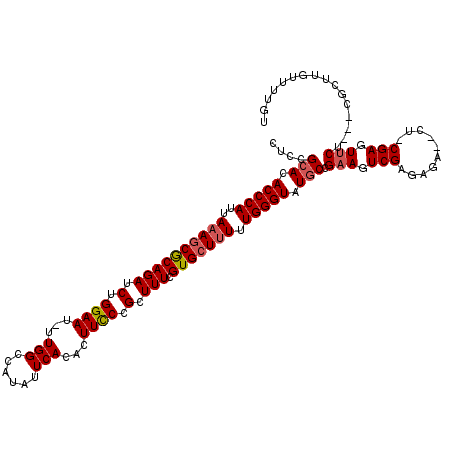

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:28 2006