
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,118,456 – 15,118,650 |
| Length | 194 |
| Max. P | 0.682194 |

| Location | 15,118,456 – 15,118,575 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.45 |
| Mean single sequence MFE | -29.53 |
| Consensus MFE | -21.25 |
| Energy contribution | -21.75 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
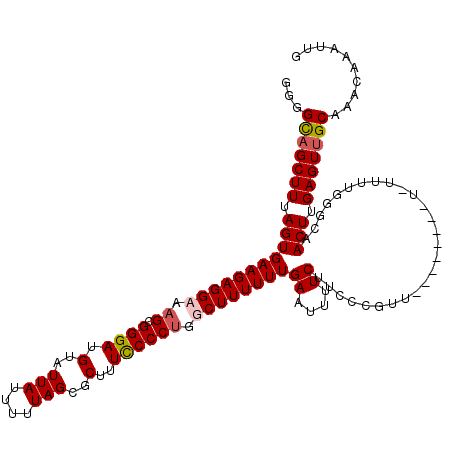
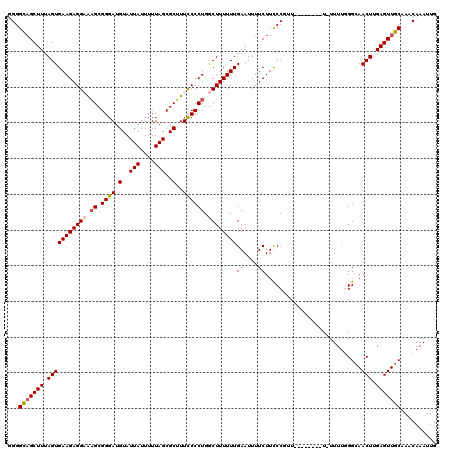
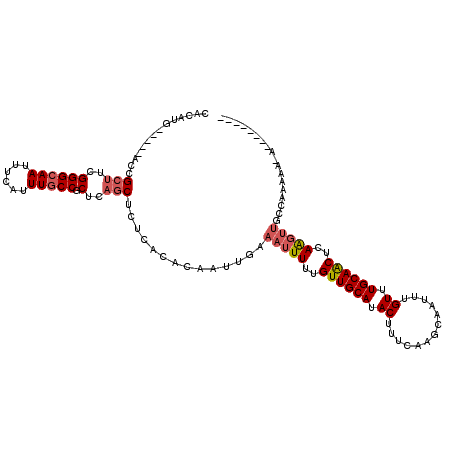
>X_DroMel_CAF1 15118456 119 + 22224390 GGGGCAGCUUUAGUGAAGAGGAAAGCGGGAUGUAUUAUUUUUAGCGCUUUUCCCUGACUUUUUUGAAUUUUCUUCCCGUUUUUUUUUUU-UUUUUGGCAACUUGAGUUGCAAACAAAUUG ...(((((((.(((((((((((((((((((.(.(((......((((........)).))......)))...).))))))))))))))))-.........))).))))))).......... ( -27.60) >DroSec_CAF1 30183 120 + 1 GGGGCAGCUUUAGUGAAGAGGAACGCGGGAUGUAUUAUUUUUAGCGCGUUUCCCUGGCUUUUUUGAAUUUUCUUCCCGUUUUUUUCUUUUUUUGUGGCAACUUGAGUUGCAAACAAAUUG ...(((((((.((((((((((((.((((((.(.(((......(((.((......)))))......)))...).)))))).)))))))))..........))).))))))).......... ( -30.50) >DroEre_CAF1 37314 109 + 1 UGGGUAGCUUUAGUGAAGAGGCAAGCGGGAUGUAUUAUUUUUAGCGCUUUCCCCUGGCUUUUUUGAAUUUUCUUUCC--U--------U-UUUUGGGCAACUUGAGUUGCAAACAAAUUG ((.(((((((.(((((((((((.((.((((.(..(((....)))..)..)))))).))))))))..........(((--.--------.-....)))..))).)))))))...))..... ( -28.80) >DroYak_CAF1 37541 111 + 1 UGGGUUGCUUUAGUGAAGAGGCGAGCGGGAUGUAUUAUUUUUAGCGCUUUCCCCUGGCUUUUUUGAAUUUUCUUUCCAUU--------U-UUUUGGGAAACUUGAGUUGCAAACAAAUUG ((.((.((((.(((((((((((.((.((((.(..(((....)))..)..)))))).))))))))........((((((..--------.-...))))))))).)))).))...))..... ( -31.20) >consensus GGGGCAGCUUUAGUGAAGAGGAAAGCGGGAUGUAUUAUUUUUAGCGCUUUCCCCUGGCUUUUUUGAAUUUUCUUCCCGUU________U_UUUUGGGCAACUUGAGUUGCAAACAAAUUG ...(((((((.(((((((((((.((.((((.(..(((....)))..)..)))))).))))))))((....))...........................))).))))))).......... (-21.25 = -21.75 + 0.50)

| Location | 15,118,456 – 15,118,575 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.45 |
| Mean single sequence MFE | -18.65 |
| Consensus MFE | -14.53 |
| Energy contribution | -15.65 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.78 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.523048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
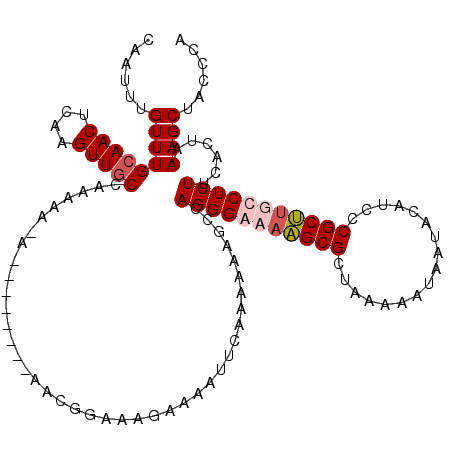
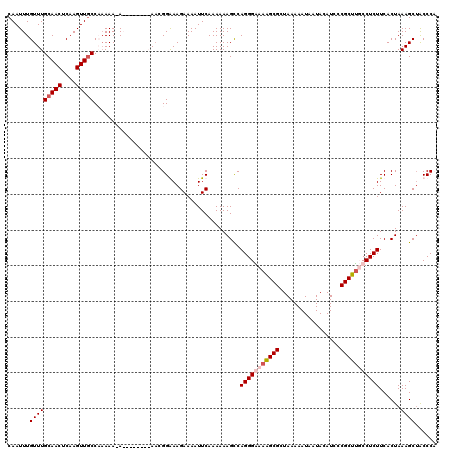
>X_DroMel_CAF1 15118456 119 - 22224390 CAAUUUGUUUGCAACUCAAGUUGCCAAAAA-AAAAAAAAAAACGGGAAGAAAAUUCAAAAAAGUCAGGGAAAAGCGCUAAAAAUAAUACAUCCCGCUUUCCUCUUCACUAAAGCUGCCCC ...((((...(((((....)))))))))..-............(((..((....)).....(((.((((.((((((.................)))))).))))..))).......))). ( -18.73) >DroSec_CAF1 30183 120 - 1 CAAUUUGUUUGCAACUCAAGUUGCCACAAAAAAAGAAAAAAACGGGAAGAAAAUUCAAAAAAGCCAGGGAAACGCGCUAAAAAUAAUACAUCCCGCGUUCCUCUUCACUAAAGCUGCCCC ...(((((..(((((....))))).)))))..((((..((..(((((.((....)).....(((...(....)..)))............)))))..))..))))............... ( -21.70) >DroEre_CAF1 37314 109 - 1 CAAUUUGUUUGCAACUCAAGUUGCCCAAAA-A--------A--GGAAAGAAAAUUCAAAAAAGCCAGGGGAAAGCGCUAAAAAUAAUACAUCCCGCUUGCCUCUUCACUAAAGCUACCCA ...((((...(((((....))))).)))).-.--------.--((...((....)).........(((((.(((((.................))))).))))).............)). ( -17.73) >DroYak_CAF1 37541 111 - 1 CAAUUUGUUUGCAACUCAAGUUUCCCAAAA-A--------AAUGGAAAGAAAAUUCAAAAAAGCCAGGGGAAAGCGCUAAAAAUAAUACAUCCCGCUCGCCUCUUCACUAAAGCAACCCA ........((((........(((((.....-.--------...))))).................(((((..((((.................))))..)))))........)))).... ( -16.43) >consensus CAAUUUGUUUGCAACUCAAGUUGCCAAAAA_A________AACGGAAAGAAAAUUCAAAAAAGCCAGGGAAAAGCGCUAAAAAUAAUACAUCCCGCUUGCCUCUUCACUAAAGCUACCCA ......(((((((((....))))).........................................(((((((((((.................)))))))))))......))))...... (-14.53 = -15.65 + 1.13)

| Location | 15,118,536 – 15,118,650 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.12 |
| Mean single sequence MFE | -23.14 |
| Consensus MFE | -14.74 |
| Energy contribution | -15.62 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562007 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15118536 114 - 22224390 CACAUG-----ACCGCUUCGGGCAAUUUCAUUUGCCGCUCAGCUCUCACACAAUUGAAAUUUUUGUUGCAUACUUUCAAGCAAUUUGUUUGCAACUCAAGUUGCCAAAAA-AAAAAAAAA ....((-----(..(((..((((((......))))).)..)))..))).....(((.(((((..((((((.((.............)).))))))..)))))..)))...-......... ( -22.72) >DroSec_CAF1 30263 115 - 1 CACAUG-----ACCGCUUCGGGCAAUUUCAUUUGCCGCUCAGCUCUCACACAAUUGAAAUUUUUGUUGCAUACUUUCAAGCAAUUUGUUUGCAACUCAAGUUGCCACAAAAAAAGAAAAA ....((-----(..(((..((((((......))))).)..)))..)))...........(((((((.(((.((((....((((.....)))).....))))))).)))))))........ ( -24.80) >DroEre_CAF1 37392 106 - 1 CACAUG-----ACCGCUUCGGGCAAUUUCAUUUGCCGCUCAACUCUCAUACAAUUGAAAUUUUUGUUGCAUACUUUCAAGCAAUUUGUUUGCAACUCAAGUUGCCCAAAA-A-------- ......-----........(((((((((.................(((......))).......((((((.((.............)).))))))..)))))))))....-.-------- ( -21.62) >DroYak_CAF1 37621 106 - 1 CACAUG-----ACCGCUUCGGGCAAUUUCAUUUGCCGCUCAGCUCUCACACAAUUGAAAUUUUUGUUGCAUACUUUCAAGCAAUUUGUUUGCAACUCAAGUUUCCCAAAA-A-------- ....((-----(..(((..((((((......))))).)..)))..))).......(((((((..((((((.((.............)).))))))..)))))))......-.-------- ( -24.62) >DroPer_CAF1 84854 102 - 1 CACAGGGAGGGGGGGCCUUGGGCAAUUUCAUUUUCCGCUUAGCUCUUACACAAUUGAAAUUUUUGUUGCAUACUUUCGAGCAAUUUGUUUGCAGCUCAGC----CA-------------- ....(((((..(..(((...)))..)..)...))))(((.(((((..(((.(((((........((.....)).......))))))))..).)))).)))----..-------------- ( -21.96) >consensus CACAUG_____ACCGCUUCGGGCAAUUUCAUUUGCCGCUCAGCUCUCACACAAUUGAAAUUUUUGUUGCAUACUUUCAAGCAAUUUGUUUGCAACUCAAGUUGCCAAAAA_A________ ..............(((..((((((......))))).)..)))..............(((((..((((((.((.............)).))))))..))))).................. (-14.74 = -15.62 + 0.88)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:06 2006