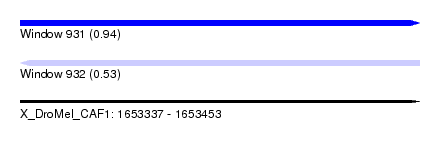

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,653,337 – 1,653,453 |
| Length | 116 |
| Max. P | 0.939469 |

| Location | 1,653,337 – 1,653,453 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 94.71 |
| Mean single sequence MFE | -36.36 |
| Consensus MFE | -28.98 |
| Energy contribution | -29.46 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939469 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1653337 116 + 22224390 UUUUAGCCCGCUUUCAGCGCGGUCUUAUCUUCCACGCAUAUUAUUAAUGCCUUGUCAUUAGGCGACUACUGUAUACCAUUA---UACAGGUGCAGGCGCCGACAGGAGCGGCAACAACA .....(((.((((((.((((((.........))..((((........(((((.......)))))....(((((((....))---)))))))))..)))).))...)))))))....... ( -35.60) >DroSec_CAF1 18181 116 + 1 UUUUAGCCCGCUUUCAGCGCGGUCUUAUCUUCCACGCAUAUUAUUAAUGCCUUGUCAUUAGGCGACUACUGUAUACCAUUA---UACAGGUGCAGGCGCCGACAGGAGCGGCAACAACA .....(((.((((((.((((((.........))..((((........(((((.......)))))....(((((((....))---)))))))))..)))).))...)))))))....... ( -35.60) >DroSim_CAF1 7431 116 + 1 UUUUAGCCCGCUUUCAGCGCGGUCUUAUCUUCCACGCAUAUUAUUAAUGCCUUGUCAUUAGGCGACUACUGUAUACCAUUA---UACAGGUGCAGGCGCCGACAGGAGCGGCAACAACA .....(((.((((((.((((((.........))..((((........(((((.......)))))....(((((((....))---)))))))))..)))).))...)))))))....... ( -35.60) >DroEre_CAF1 19803 116 + 1 GUUUGGACCGCUUUCAGCGCGGUCUUAUCUGCCACGCAUAUUAUUAAUGCCUUGUCAUUAGGCGACUACUGUAUGCCAUUA---UACAGGUGCAGGCGCCGACAGGAGCGGCAGCAACA (((.(((((((.......)))))))...(((((.(((((.......))))((((((....((((((....)).))))....---....((((....)))))))))).).))))).))). ( -42.80) >DroYak_CAF1 18906 115 + 1 UUUUAGCCCGUUUUCAGCGCGGUCUUAUCUGCCACGCAUAUUAUUAAUGCCUUGUCAUUAGGCGACUACUGUAUACCAUUAUAAUACAGGUGCAGGCGCCGAAAGGAG----AACAACA .........((((((.(((((((.......)))..((((........(((((.......)))))....((((((.........))))))))))..))))(....))))----))).... ( -32.20) >consensus UUUUAGCCCGCUUUCAGCGCGGUCUUAUCUUCCACGCAUAUUAUUAAUGCCUUGUCAUUAGGCGACUACUGUAUACCAUUA___UACAGGUGCAGGCGCCGACAGGAGCGGCAACAACA .......(((((....(((.((.........)).))).............((((((....((((....((((..(((...........))))))).)))))))))))))))........ (-28.98 = -29.46 + 0.48)

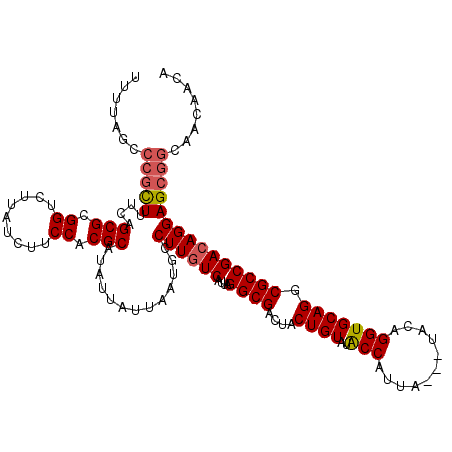
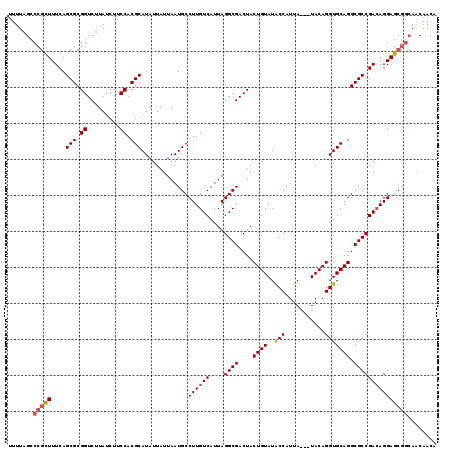
| Location | 1,653,337 – 1,653,453 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 94.71 |
| Mean single sequence MFE | -38.02 |
| Consensus MFE | -28.24 |
| Energy contribution | -29.64 |
| Covariance contribution | 1.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.533283 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1653337 116 - 22224390 UGUUGUUGCCGCUCCUGUCGGCGCCUGCACCUGUA---UAAUGGUAUACAGUAGUCGCCUAAUGACAAGGCAUUAAUAAUAUGCGUGGAAGAUAAGACCGCGCUGAAAGCGGGCUAAAA .......(((((....).))))((((((..(((((---((....)))))))...((((((.......))))...........((((((.........)))))).))..))))))..... ( -39.20) >DroSec_CAF1 18181 116 - 1 UGUUGUUGCCGCUCCUGUCGGCGCCUGCACCUGUA---UAAUGGUAUACAGUAGUCGCCUAAUGACAAGGCAUUAAUAAUAUGCGUGGAAGAUAAGACCGCGCUGAAAGCGGGCUAAAA .......(((((....).))))((((((..(((((---((....)))))))...((((((.......))))...........((((((.........)))))).))..))))))..... ( -39.20) >DroSim_CAF1 7431 116 - 1 UGUUGUUGCCGCUCCUGUCGGCGCCUGCACCUGUA---UAAUGGUAUACAGUAGUCGCCUAAUGACAAGGCAUUAAUAAUAUGCGUGGAAGAUAAGACCGCGCUGAAAGCGGGCUAAAA .......(((((....).))))((((((..(((((---((....)))))))...((((((.......))))...........((((((.........)))))).))..))))))..... ( -39.20) >DroEre_CAF1 19803 116 - 1 UGUUGCUGCCGCUCCUGUCGGCGCCUGCACCUGUA---UAAUGGCAUACAGUAGUCGCCUAAUGACAAGGCAUUAAUAAUAUGCGUGGCAGAUAAGACCGCGCUGAAAGCGGUCCAAAC .....(((((((...((((((((.((((...((((---(......))))))))).))))....))))..((((.......)))))))))))....((((((.......))))))..... ( -42.90) >DroYak_CAF1 18906 115 - 1 UGUUGUU----CUCCUUUCGGCGCCUGCACCUGUAUUAUAAUGGUAUACAGUAGUCGCCUAAUGACAAGGCAUUAAUAAUAUGCGUGGCAGAUAAGACCGCGCUGAAAACGGGCUAAAA ....(((----(...(((((((((((((((.((((((((..((.....)).((((.((((.......)))))))))))))))).)).))))........)))))))))..))))..... ( -29.60) >consensus UGUUGUUGCCGCUCCUGUCGGCGCCUGCACCUGUA___UAAUGGUAUACAGUAGUCGCCUAAUGACAAGGCAUUAAUAAUAUGCGUGGAAGAUAAGACCGCGCUGAAAGCGGGCUAAAA .......(((((....).))))((((((..(((((...........)))))...((((((.......))))...........((((((.........)))))).))..))))))..... (-28.24 = -29.64 + 1.40)

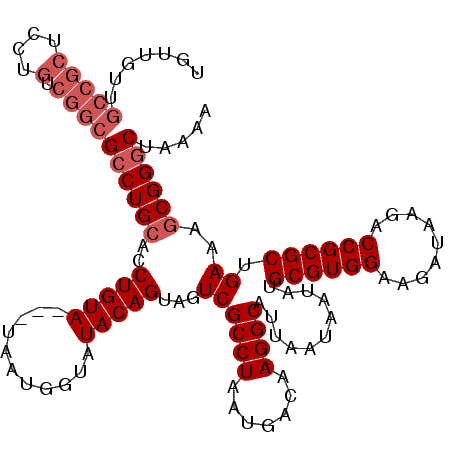
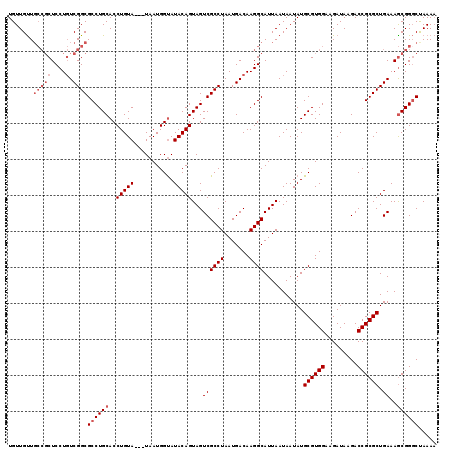
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:48:33 2006