
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,064,403 – 15,064,562 |
| Length | 159 |
| Max. P | 0.697272 |

| Location | 15,064,403 – 15,064,522 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 88.18 |
| Mean single sequence MFE | -44.40 |
| Consensus MFE | -32.21 |
| Energy contribution | -33.90 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.697272 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15064403 119 - 22224390 AGGUGGACUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCUGUGUUGUGGGGAGGGUGUCUUGGUAGAGUCCCGUAUCCGGACAUUGAUGCACCCGGAUGAGCCGGAUUCG .(((((((((((((....(((((..((...((...((((.....))))...))...))..))))).....))))))))))..(((((((............))))))).)))....... ( -41.80) >DroSec_CAF1 25658 115 - 1 AGGUGGACUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCUG----GAGGGGGCGGGGUCUUGGUAGAGUCCCGGAUCCGGGCAUUGAUGCACCCGGAUGAGCCGGAUUCG ....((((((((((...((((((..(((((.((..((((.....)))).)----))))))..))))))..))))))))))(((((((((.((((..........))))..))))))))) ( -49.50) >DroSim_CAF1 12056 115 - 1 AGGUGGACUUUGCUUUUGGACCUUACUUUUGCCUUGGCUUGAUUAGUCUG----GAGGGGGCGGGGUCUUGGUAGAGUCCCGGAUCCGGACAUUGAUGCACCCGGAUGAGCCGGCUUCG ((((((((((((((...((((((..((((..((..((((.....)))).)----)..))))..)))))).))))))))))(((((((((............))))))...))))))).. ( -47.10) >DroEre_CAF1 16702 106 - 1 AGGUCGGCUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCCG------------GGGGUCUUGGUGGAGUCCCGGAUCCGGACAUUGAUGCACCCGGAUGAGUCGAGUGC- .....((((........)))).........((((((((((......((((------------(((.(((....))).)))))))(((((............))))).)))))))).))- ( -39.20) >consensus AGGUGGACUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCUG____GAGGGGGCGGGGUCUUGGUAGAGUCCCGGAUCCGGACAUUGAUGCACCCGGAUGAGCCGGAUUCG .(((((((((((((...((((((..(((((.....((((.....))))......)))))...))))))..))))))))))...((((((............))))))..)))....... (-32.21 = -33.90 + 1.69)



| Location | 15,064,443 – 15,064,562 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 86.51 |
| Mean single sequence MFE | -39.50 |
| Consensus MFE | -29.50 |
| Energy contribution | -30.50 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.605055 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15064443 119 - 22224390 GACCCCCCCCCCCCUCACUCCUCACUUGCACCUGUAGCCCAGGUGGACUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCUGUGUUGUGGGGAGGGUGUCUUGGUAGAGUC ((((((.((((.(..(((..........((((((.....))))))((((..(((...(((..........)))..)))......)))).)))..).)))).))).)))........... ( -39.60) >DroSec_CAF1 25698 103 - 1 ------------CCUCACUUCUCACGUGCACCUGUAGCCCAGGUGGACUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCUG----GAGGGGGCGGGGUCUUGGUAGAGUC ------------...(((.......)))((((((.....))))))(((((((((...((((((..(((((.((..((((.....)))).)----))))))..))))))..))))))))) ( -38.90) >DroSim_CAF1 12096 103 - 1 ------------CCUCACUCCUCACUUGCACCUGUAGCCCAGGUGGACUUUGCUUUUGGACCUUACUUUUGCCUUGGCUUGAUUAGUCUG----GAGGGGGCGGGGUCUUGGUAGAGUC ------------................((((((.....))))))(((((((((...((((((..((((..((..((((.....)))).)----)..))))..)))))).))))))))) ( -40.00) >consensus ____________CCUCACUCCUCACUUGCACCUGUAGCCCAGGUGGACUUUGCUUUUGGCCCUUACUUUUGCCUUGGCUUGAUUAGUCUG____GAGGGGGCGGGGUCUUGGUAGAGUC ............................((((((.....))))))(((((((((...((((((..(((((.....((((.....))))......)))))...))))))..))))))))) (-29.50 = -30.50 + 1.00)

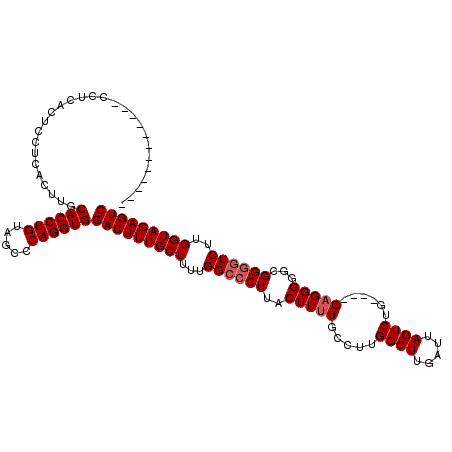

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:57:05 2006