
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,048,306 – 15,048,405 |
| Length | 99 |
| Max. P | 0.980589 |

| Location | 15,048,306 – 15,048,405 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 82.94 |
| Mean single sequence MFE | -26.60 |
| Consensus MFE | -20.22 |
| Energy contribution | -19.67 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.87 |
| SVM RNA-class probability | 0.980589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15048306 99 + 22224390 GGGCGGGGAAAAUUGUUGCCGCUUUCAGUGGUUGAAAAAAAA--AGAAUCCGAGAAGGAAAAAAAAACUGCGAGCAGAGAAAACUCAGCGGAA-AACCCGCU .((((((........((((.((((.((((..((....))...--....(((.....))).......)))).)))).(((....))).))))..-..)))))) ( -26.40) >DroSec_CAF1 4222 94 + 1 AAGCGGGGAAAAUUGUUGCCGCUUUUAGUGGUUUAAAAAAAA---AACCCCGAGAAGGAA----AAACUGCGAGCAGAGAAAACUCAGCGGAA-AACCCGCU .((((((........((((.((((.(((((((((.......)---))))((.....))..----..)))).)))).(((....))).))))..-..)))))) ( -27.00) >DroEre_CAF1 3991 97 + 1 GGGCGGGGAAAACUGUUGCCGUUUUCAGUGGUUUAAAGAAGAAAAAAAGCGGAGUAGGA-----AAGCUGCGAGCAGUGAAAACUCAGCGGAAAAACGCGCU .((((((....((((((((.((((((..(.((((............)))).).....))-----)))).)).)))))).....))).(((......)))))) ( -26.40) >consensus GGGCGGGGAAAAUUGUUGCCGCUUUCAGUGGUUUAAAAAAAA__AAAACCCGAGAAGGAA____AAACUGCGAGCAGAGAAAACUCAGCGGAA_AACCCGCU .((((((.....((...(((((.....)))))....)).............................(((((((.........))).)))).....)))))) (-20.22 = -19.67 + -0.55)


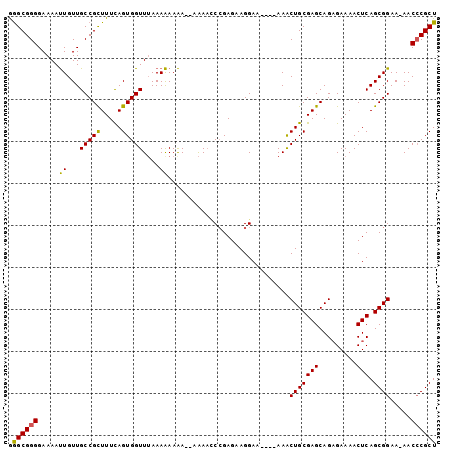
| Location | 15,048,306 – 15,048,405 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 82.94 |
| Mean single sequence MFE | -22.23 |
| Consensus MFE | -14.50 |
| Energy contribution | -14.83 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.883025 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15048306 99 - 22224390 AGCGGGUU-UUCCGCUGAGUUUUCUCUGCUCGCAGUUUUUUUUUCCUUCUCGGAUUCU--UUUUUUUUCAACCACUGAAAGCGGCAACAAUUUUCCCCGCCC .(((((..-.......(((....)))(((.(((..........(((.....)))....--.....(((((.....))))))))))).........))))).. ( -23.30) >DroSec_CAF1 4222 94 - 1 AGCGGGUU-UUCCGCUGAGUUUUCUCUGCUCGCAGUUU----UUCCUUCUCGGGGUU---UUUUUUUUAAACCACUAAAAGCGGCAACAAUUUUCCCCGCUU ((((((..-....((.(((....))).))..((.((((----((((.....))((((---(.......)))))...)))))).))..........)))))). ( -25.70) >DroEre_CAF1 3991 97 - 1 AGCGCGUUUUUCCGCUGAGUUUUCACUGCUCGCAGCUU-----UCCUACUCCGCUUUUUUUCUUCUUUAAACCACUGAAAACGGCAACAGUUUUCCCCGCCC ...(((.......((.((((.....(((....)))...-----....)))).))......................(((((((....).))))))..))).. ( -17.70) >consensus AGCGGGUU_UUCCGCUGAGUUUUCUCUGCUCGCAGUUU____UUCCUUCUCGGAUUUU__UUUUUUUUAAACCACUGAAAGCGGCAACAAUUUUCCCCGCCC .(((((.......((.((((.......)))))).................................................(....).......))))).. (-14.50 = -14.83 + 0.33)

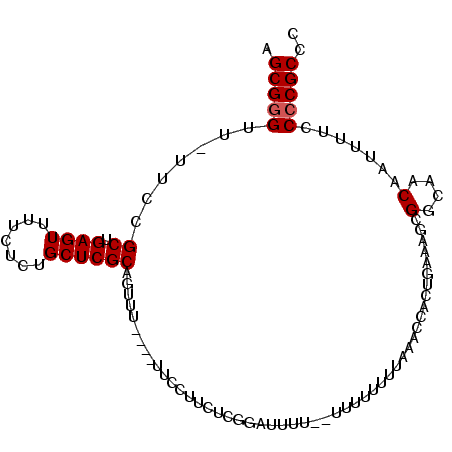
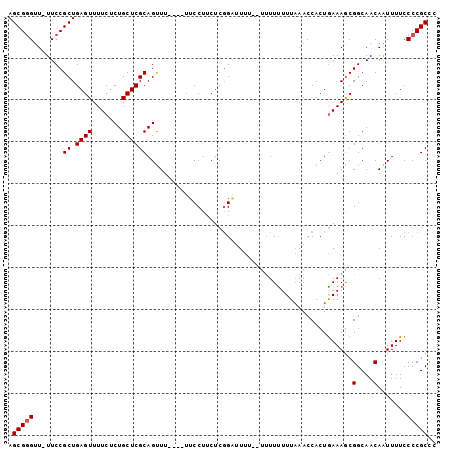
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:46 2006