
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,045,921 – 15,046,060 |
| Length | 139 |
| Max. P | 0.881158 |

| Location | 15,045,921 – 15,046,020 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.34 |
| Mean single sequence MFE | -31.98 |
| Consensus MFE | -20.50 |
| Energy contribution | -20.62 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.64 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.525745 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15045921 99 - 22224390 AGUUUCUGAUUGCGAAACACAAAAAGGGA--G--CCAAAAUGGUAGGAAAACAUGGGGAGCUGGCUGACAGCGGACCUA-----------------GUCGAUGUCGAUGAUGGUUCCCUG .(((((.......)))))......(((((--(--(((................((((..((((.....))))...))))-----------------((((....))))..))))))))). ( -34.40) >DroSec_CAF1 1712 97 - 1 AGUUUCUGAUUGCGAAGCACAAAAAGGGA--G--CAAAAAUGGAAGGAGAACAUGGGGAGCUGGCUGACAGCGGACCUA-----------------GUCGAUGUCGAUGAUGG--CCCUG .(((((.......))))).......(((.--(--(....((.((....((...((((..((((.....))))...))))-----------------.))....)).))....)--)))). ( -26.70) >DroEre_CAF1 1477 97 - 1 AGUUUCUGAUUGCAAAGCACAAAAAGGGG-----GCAAAAUGGUAGGAAAACAUGGGAAGCUGGCUGACAGCGGACCCGU------------------CGAUGUCGAUGAUGGUUCCCUG .((((.((....))))))......(((((-----((...((.((.((.....(((((..((((.....))))...)))))------------------.....)).)).)).))))))). ( -26.40) >DroYak_CAF1 1527 120 - 1 AGUUUCUGAUUGCAAAGCACAAAAGGGGGGAGCGGCAAAAUGGUAGGAUAACUUGGGAAGCUGGCUGACAGCGGACCCGUCCAUGUCGAUGUCGAUGUCGAUGUCGAUGAUGGUUCCCUG ....(((...(((...)))....)))((((((((((.....(((......))).(((..((((.....))))...))))))(((((((((((((....)))))))))).)))))))))). ( -40.40) >consensus AGUUUCUGAUUGCAAAGCACAAAAAGGGA__G__CCAAAAUGGUAGGAAAACAUGGGAAGCUGGCUGACAGCGGACCCA_________________GUCGAUGUCGAUGAUGGUUCCCUG .(((((.......)))))......((((.........................((((..((((.....))))...)))).................((((....)))).......)))). (-20.50 = -20.62 + 0.12)

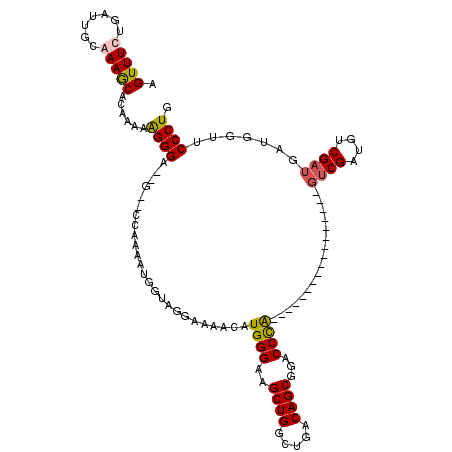

| Location | 15,045,945 – 15,046,060 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.54 |
| Mean single sequence MFE | -26.63 |
| Consensus MFE | -22.99 |
| Energy contribution | -22.55 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.881158 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15045945 115 - 22224390 AAAAAGGAACAACGGAAAAAAUCAUCAAUAGGCCAUGAUGAGUUUCUGAUUGCGAAACACAAAAAGGGA--G--CCAAAAUGGUAGGAAAACAUGGGGAGCUGGCUGACAGCGGACCUA- .........(((((((((...((((((........)))))).)))))).))).................--(--((.....))).........((((..((((.....))))...))))- ( -25.90) >DroSec_CAF1 1734 113 - 1 AAA--AGAACAACGGAAAAAAUCAUCAAUAGGCCAUGAUGAGUUUCUGAUUGCGAAGCACAAAAAGGGA--G--CAAAAAUGGAAGGAGAACAUGGGGAGCUGGCUGACAGCGGACCUA- ...--.......((((((...((((((........)))))).)))))).((((................--)--)))................((((..((((.....))))...))))- ( -24.89) >DroEre_CAF1 1499 115 - 1 AAAAAAGAACAACGGGUAAAAUCAUCAAUAGGCCAUGAUGAGUUUCUGAUUGCAAAGCACAAAAAGGGG-----GCAAAAUGGUAGGAAAACAUGGGAAGCUGGCUGACAGCGGACCCGU ...........((((((....((((......(((((..((..(..((..(((.......)))..))..)-----.))..)))))........))))...((((.....))))..)))))) ( -26.54) >DroYak_CAF1 1567 120 - 1 AAAAAAGAAGAACGGAAAAAAUCAUCAAUAGGCCAUGAUGAGUUUCUGAUUGCAAAGCACAAAAGGGGGGAGCGGCAAAAUGGUAGGAUAACUUGGGAAGCUGGCUGACAGCGGACCCGU .......(((..((((((...((((((........)))))).)))))).((((...((.(........)..)).)))).............)))(((..((((.....))))...))).. ( -29.20) >consensus AAAAAAGAACAACGGAAAAAAUCAUCAAUAGGCCAUGAUGAGUUUCUGAUUGCAAAGCACAAAAAGGGA__G__CCAAAAUGGUAGGAAAACAUGGGAAGCUGGCUGACAGCGGACCCA_ ............((((((...((((((........)))))).)))))).............................................((((..((((.....))))...)))). (-22.99 = -22.55 + -0.44)

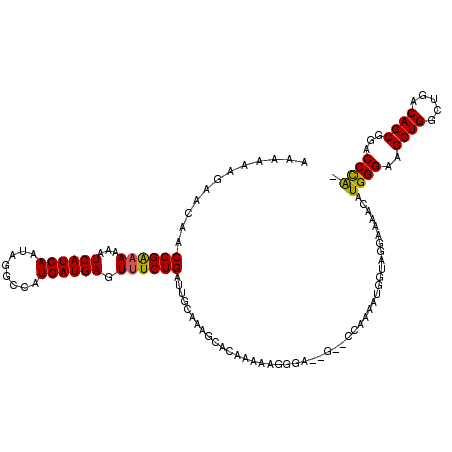

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:41 2006