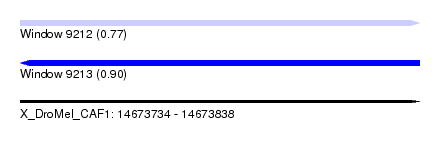

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,673,734 – 14,673,838 |
| Length | 104 |
| Max. P | 0.904861 |

| Location | 14,673,734 – 14,673,838 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 89.47 |
| Mean single sequence MFE | -30.66 |
| Consensus MFE | -21.64 |
| Energy contribution | -22.70 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.770672 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14673734 104 + 22224390 CCAAUCAGAAUGCACGCACACACUAAGGGUUAAUUUUCUAGUGUGUAUGCGUGUGCGUGUCUGUG--UGUGCGGGUUAUUUUUGUUGUUUGGUAUUUAUACACACC ......(((((((((((((((((((..((.....))..))))))))....)))))))).)))(((--((((..(((.................)))..))))))). ( -31.33) >DroSec_CAF1 6451 103 + 1 CCAAUCAGAAUGCACGCACACACUAAGGGUUAAUUUUCUAGUGUGUAUGCGUGUGCAUGGCUGUG--UGUGCGGUUUAUUUUUGUUGU-UGGUAUUUAUACAGACA (((((((((((((((((((((((((..((.....))..))))))))....))))))))((((((.--...))))))....))))..))-))).............. ( -30.40) >DroSim_CAF1 6378 105 + 1 CCAAUCAGAAUGCACGCACACACUAAGGGUUAAUUUUCUAGUGUGUAUGCGUGUGCAUGGCUGUGUGUGUGCGGUUUAUUUUUGUUGU-UGGUAUUUAUACACACA .....(((.((((((((((((((((..((.....))..))))))))....))))))))..)))((((((((..(..((((........-.)))).)..)))))))) ( -33.60) >DroEre_CAF1 7655 97 + 1 CCAAUCAGAAUGCACGCACACACUAAGGGUUAAUUUUCCAGUGUGUAUGCCUGUGUGU------G--UGUGCCUCGUAUUUUUGUUGU-UGGUAUUUAUGCACACC ((((((((((.((((((((((((...((((..((........))....))))))))))------)--))))).......)))))..))-))).............. ( -27.30) >consensus CCAAUCAGAAUGCACGCACACACUAAGGGUUAAUUUUCUAGUGUGUAUGCGUGUGCAUGGCUGUG__UGUGCGGUUUAUUUUUGUUGU_UGGUAUUUAUACACACA (((..((((((((((((((((((((.(((......)))))))))))....)))))))).(((((......))))).....)))).....))).............. (-21.64 = -22.70 + 1.06)



| Location | 14,673,734 – 14,673,838 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 89.47 |
| Mean single sequence MFE | -24.77 |
| Consensus MFE | -18.01 |
| Energy contribution | -18.82 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.904861 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14673734 104 - 22224390 GGUGUGUAUAAAUACCAAACAACAAAAAUAACCCGCACA--CACAGACACGCACACGCAUACACACUAGAAAAUUAACCCUUAGUGUGUGCGUGCAUUCUGAUUGG (((((......))))).......................--..(((((((((((((((.........................)))))))))))...))))..... ( -26.71) >DroSec_CAF1 6451 103 - 1 UGUCUGUAUAAAUACCA-ACAACAAAAAUAAACCGCACA--CACAGCCAUGCACACGCAUACACACUAGAAAAUUAACCCUUAGUGUGUGCGUGCAUUCUGAUUGG (((..(((....)))..-)))..................--..(((..((((((..((((((((...((..........))..)))))))))))))).)))..... ( -21.30) >DroSim_CAF1 6378 105 - 1 UGUGUGUAUAAAUACCA-ACAACAAAAAUAAACCGCACACACACAGCCAUGCACACGCAUACACACUAGAAAAUUAACCCUUAGUGUGUGCGUGCAUUCUGAUUGG (((((((..........-................)))))))..(((..((((((..((((((((...((..........))..)))))))))))))).)))..... ( -26.07) >DroEre_CAF1 7655 97 - 1 GGUGUGCAUAAAUACCA-ACAACAAAAAUACGAGGCACA--C------ACACACAGGCAUACACACUGGAAAAUUAACCCUUAGUGUGUGCGUGCAUUCUGAUUGG (((((......))))).-.............(((((((.--(------((((((.(.....).....((........))....))))))).)))).)))....... ( -25.00) >consensus GGUGUGUAUAAAUACCA_ACAACAAAAAUAAACCGCACA__CACAGCCACGCACACGCAUACACACUAGAAAAUUAACCCUUAGUGUGUGCGUGCAUUCUGAUUGG (((((......)))))..................((((............((....)).((((((((((...........)))))))))).))))........... (-18.01 = -18.82 + 0.81)

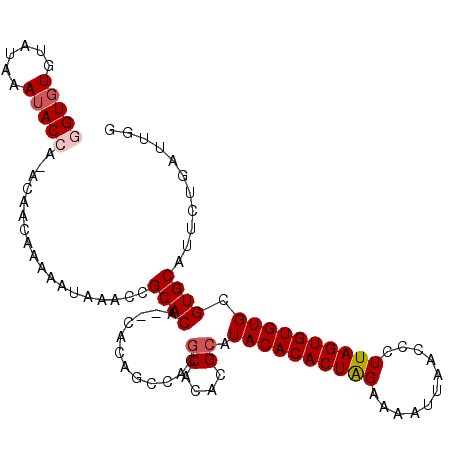

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:44 2006