
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,570,997 – 14,571,127 |
| Length | 130 |
| Max. P | 0.956365 |

| Location | 14,570,997 – 14,571,091 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.17 |
| Mean single sequence MFE | -32.39 |
| Consensus MFE | -25.63 |
| Energy contribution | -25.63 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.956365 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14570997 94 - 22224390 CUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA-------GGCAAAUAAAAGCCAUCCCAAAGAGCA----------- ...((((..(((((((...-------(((((....)))((....))...))-..)))))))....(....)))))-------(((........))).............----------- ( -27.50) >DroPse_CAF1 52596 120 - 1 GUGCGGCACGUAGUCGCCGCAGUUGCAUGUUGAAAAGCGCGAAAGCGUAGCCGUUGACUGCCAAAUGAAAAGCCAGUCCAAAGGCAAAUAAAUGCCGGCCAAAAGGGCUUUCCACUGAUG ((((((((.(((((((.((..(((((..(((....)))((....))))))))).)))))))..........(((........))).......)))))(((.....)))....)))..... ( -40.50) >DroEre_CAF1 40913 94 - 1 CUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA-------GGCAAAUAAAAGCCAUCCAAAAGAGCA----------- ...((((..(((((((...-------(((((....)))((....))...))-..)))))))....(....)))))-------(((........))).............----------- ( -27.50) >DroYak_CAF1 42942 94 - 1 CUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA-------GGCAAAUAAAAGCCAGCCAAAAGAGCA----------- ...((((..(((((((...-------(((((....)))((....))...))-..)))))))..............-------(((........))).))))........----------- ( -29.60) >DroAna_CAF1 71441 93 - 1 CUGCAGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAA-AGCCA-------GGCAAAUAAAAGCCAGCCAAAAGGGCA----------- ...((....(((((((...-------(((((....)))((....))...))-..)))))))....))..-.....-------(((........))).(((.....))).----------- ( -27.80) >DroPer_CAF1 52100 120 - 1 CUGCGGCACGUAGUCGCCGCAGUUGCAUGUUGAAAAGCGCGAAAGCGUAGCCGUUGACUGCCAAAUGAAAAGCCAGUCCAAAGGCAAAUAAAUGCCGGCCAAAAGGGCUUUCCACUGAUG (((((((........)))))))..(((.((..(...((((....)))).....)..)))))....((.((((((........((((......)))).........)))))).))...... ( -41.43) >consensus CUGCGGCACGUAGUCGUUA_______GCGUUGAAAAGCGCGAAAGCGUAGC_AUUGACUGCCAAAUGAAAAGCCA_______GGCAAAUAAAAGCCAGCCAAAAGAGCA___________ ...((((..((((((.............(((....)))((....)).........))))))..........(((........)))........))))....................... (-25.63 = -25.63 + 0.00)

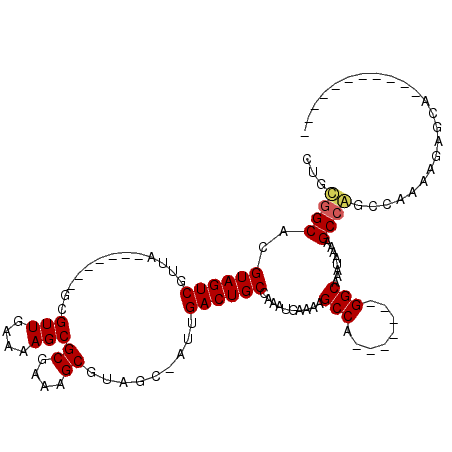

| Location | 14,571,024 – 14,571,127 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 83.41 |
| Mean single sequence MFE | -32.74 |
| Consensus MFE | -25.61 |
| Energy contribution | -25.58 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.38 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.939109 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14571024 103 - 22224390 UAUUAAAGAGGGAAU-CUUGGUAAAAGGCAGUUAA-AACUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA----- .....((((.....)-)))(((....(((((((((-...(((...((((..((....-------(((((....)))))))..)))).))-))))))))))...(....)))).----- ( -28.90) >DroSec_CAF1 46674 93 - 1 ----------GGGGU-CUUGGUAAAAGGCAGUUAA-AACUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA----- ----------..((.-(((..((...(((((((((-...(((...((((..((....-------(((((....)))))))..)))).))-))))))))))...))..))))).----- ( -29.70) >DroEre_CAF1 40940 102 - 1 CAUUAAAGAGGG-GU-CUUGGUAAAAGGCAGUUAA-AACUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA----- ...........(-(.-(((..((...(((((((((-...(((...((((..((....-------(((((....)))))))..)))).))-))))))))))...))..))))).----- ( -29.70) >DroYak_CAF1 42969 103 - 1 CAUUAAAGAGGG-UUCCUUGGUAAAAGGCAGUUAA-AACUGUGGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAAAAGCCA----- ..(((..((((.-..))))..)))...(((((...-.)))))(((..(((((((...-------(((((....)))((....))...))-..)))))))....(....)))).----- ( -29.60) >DroAna_CAF1 71468 93 - 1 ---------GGGGCU-GUAGGUAAAAGGCAGUUAA-AACUGCAGCACGUAGUCGUUA-------GCGUUGAAAAGCGCGAAAGCGUAGC-AUUGACUGCCAAAUGAA-AGCCA----- ---------..((((-..........(((((((((-..((((.(((((....)))..-------(((((....)))))....)))))).-.))))))))).......-)))).----- ( -34.13) >DroPer_CAF1 52140 107 - 1 ----------GGCAG-AGUGGUAAAAGGCAGUUAAAAACUGCGGCACGUAGUCGCCGCAGUUGCAUGUUGAAAAGCGCGAAAGCGUAGCCGUUGACUGCCAAAUGAAAAGCCAGUCCA ----------....(-(.((((....(((((((((.(((((((((........)))))))))((..(((....)))((....))...))..)))))))))...(....))))).)).. ( -44.40) >consensus _________GGG_AU_CUUGGUAAAAGGCAGUUAA_AACUGUGGCACGUAGUCGUUA_______GCGUUGAAAAGCGCGAAAGCGUAGC_AUUGACUGCCAAAUGAAAAGCCA_____ ...................(((....(((((((((...((((.(((((....))).........(((((....)))))....))))))...)))))))))...(....))))...... (-25.61 = -25.58 + -0.03)

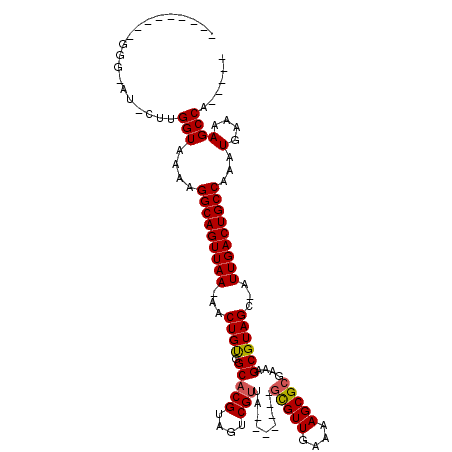

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:52:55 2006