
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,570,392 – 14,570,500 |
| Length | 108 |
| Max. P | 0.987093 |

| Location | 14,570,392 – 14,570,500 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 73.96 |
| Mean single sequence MFE | -30.48 |
| Consensus MFE | -13.41 |
| Energy contribution | -15.73 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.19 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.44 |
| SVM decision value | 2.07 |
| SVM RNA-class probability | 0.987093 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14570392 108 + 22224390 UGCAUUCCUCAUUAAAUUGGAGUCUGUAUGAAAGCCCACACCCACCCAUGUGUA--UGUAUG-----UG--UAUGUAUAUAGAAUCUGUGUGGAUAUAGCCAC--ACAUGCAGGCUUGC .(((..((((((......(((.((((((((...((..(((..(((....)))..--)))..)-----).--....)))))))).)))((((((......))))--))))).)))..))) ( -31.30) >DroSec_CAF1 46055 100 + 1 UGCAUUCCUCAUUAAAUUGGAGUCUGUAUGAAAGCGCACACCCAACGAUGUG-------------------UGUGUAUAUAACAUCUGUGUAGAUAUAGCCACACACAUGCAGGCUAUC .....(((..........)))(((((((((...((((((((........)))-------------------)))))..........(((((.(......).)))))))))))))).... ( -29.20) >DroEre_CAF1 40341 113 + 1 UGCAUUCCUCAUUAAAUUGGAGUCUGCAUGAAAGCUCACACCCACCGAUGUGUACAUAUAUAGUGUGUAUAUAGCUAUGUAAUAUCUGUGUGGAUAUAUGCACACA------UGCUUGC ((((..(..((......))..)..))))...((((..((((........)))).........(((((((((((.(((..((.....))..))).))))))))))).------.)))).. ( -30.70) >DroYak_CAF1 42362 110 + 1 UGCAUUCCUCAUUAAAUUGGAGUCUGCAUGGAAACCCACACCCACA----UGUACAUACAUA-----UAUAUGUAUAUAUAAUAUCUUGGUAGAUAUAUUCACACACAUGCAGGCUCGC ...................(((((((((((...............(----((((((((....-----..)))))))))...((((((....)))))).........))))))))))).. ( -30.70) >consensus UGCAUUCCUCAUUAAAUUGGAGUCUGCAUGAAAGCCCACACCCACCGAUGUGUA__UAUAUA_____UA__UAUGUAUAUAAUAUCUGUGUAGAUAUAGCCACACACAUGCAGGCUUGC ...................(((((((((((.......((((........))))............................((((((....)))))).........))))))))))).. (-13.41 = -15.73 + 2.31)

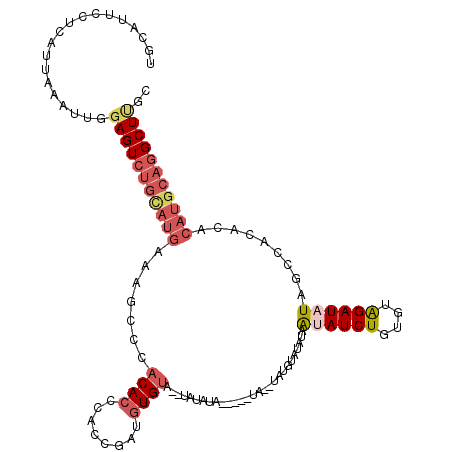

| Location | 14,570,392 – 14,570,500 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 73.96 |
| Mean single sequence MFE | -29.89 |
| Consensus MFE | -14.06 |
| Energy contribution | -18.00 |
| Covariance contribution | 3.94 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.47 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.837319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14570392 108 - 22224390 GCAAGCCUGCAUGU--GUGGCUAUAUCCACACAGAUUCUAUAUACAUA--CA-----CAUACA--UACACAUGGGUGGGUGUGGGCUUUCAUACAGACUCCAAUUUAAUGAGGAAUGCA (((((((..(((((--((((......))))))................--((-----(...((--(....))).))).)))..))))...........(((..........))).))). ( -29.10) >DroSec_CAF1 46055 100 - 1 GAUAGCCUGCAUGUGUGUGGCUAUAUCUACACAGAUGUUAUAUACACA-------------------CACAUCGUUGGGUGUGCGCUUUCAUACAGACUCCAAUUUAAUGAGGAAUGCA .(((.((.(((((((((((...((((((....))))))......))))-------------------))))).)).)).)))(((.((((((...............))))))..))). ( -25.96) >DroEre_CAF1 40341 113 - 1 GCAAGCA------UGUGUGCAUAUAUCCACACAGAUAUUACAUAGCUAUAUACACACUAUAUAUGUACACAUCGGUGGGUGUGAGCUUUCAUGCAGACUCCAAUUUAAUGAGGAAUGCA (((.(((------((...((.(((((((((...(((..((((((..((((.......))))))))))...))).))))))))).))...)))))....(((..........))).))). ( -32.10) >DroYak_CAF1 42362 110 - 1 GCGAGCCUGCAUGUGUGUGAAUAUAUCUACCAAGAUAUUAUAUAUACAUAUA-----UAUGUAUGUACA----UGUGGGUGUGGGUUUCCAUGCAGACUCCAAUUUAAUGAGGAAUGCA (.(((((..(((.((..((...((((((....))))))..(((((((((...-----.)))))))))))----..)).)))..))))).).((((..(((.........)))...)))) ( -32.40) >consensus GCAAGCCUGCAUGUGUGUGGAUAUAUCCACACAGAUAUUAUAUACACA__UA_____UAUAUA__UACACAUCGGUGGGUGUGGGCUUUCAUACAGACUCCAAUUUAAUGAGGAAUGCA (.(((((((((((((((((...(((((......)))))..)))))))....................(((....))).)))))))))).).((((..(((.........)))...)))) (-14.06 = -18.00 + 3.94)

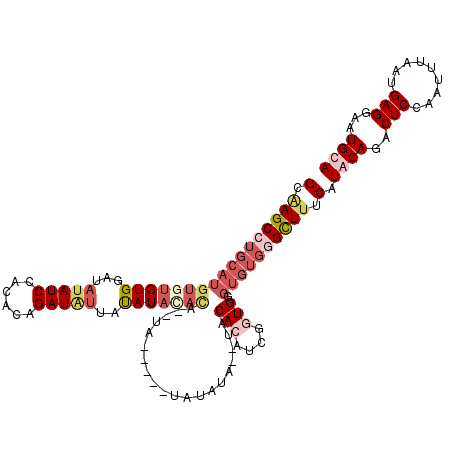
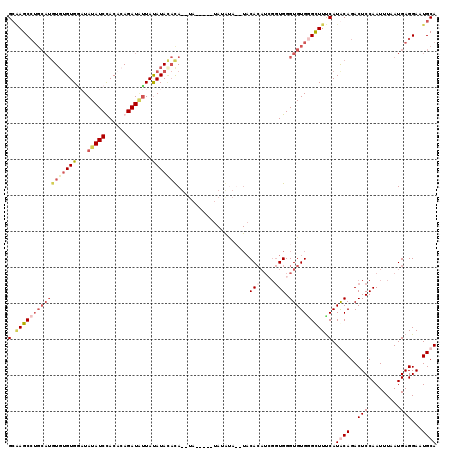
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:52:52 2006