
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,478,396 – 14,478,562 |
| Length | 166 |
| Max. P | 0.994829 |

| Location | 14,478,396 – 14,478,505 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.12 |
| Mean single sequence MFE | -29.16 |
| Consensus MFE | -28.45 |
| Energy contribution | -28.05 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.98 |
| SVM decision value | 2.51 |
| SVM RNA-class probability | 0.994829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14478396 109 - 22224390 GGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA-ACUC---GCAACAACUUCUCAGUUCCUCAUCUGCGACUGC-------AACUGCAUCUUCAGCUCC ((((....(((((((.((.(.((((......))))).)).)))))))..)))).-....---((.......((.(((........))).)).(((-------....)))......))... ( -29.70) >DroSec_CAF1 3599 109 - 1 GGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA-ACUC---GCAACAACUUCUCAGUUCCUCAUCUGCAACUGC-------AACUGCAUCUUUAGCUCC ((((....(((((((.((.(.((((......))))).)).)))))))..)))).-....---(((..((((....)))).......(((....))-------)..)))............ ( -27.90) >DroSim_CAF1 10099 109 - 1 GGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA-ACUC---GCAACAACUUCUCAGUUCCUCAUCUGCAACUGC-------AUCUGCAUCUUCAGCUCC ((((....(((((((.((.(.((((......))))).)).)))))))..)))).-....---(((..((((....)))).......(((....))-------)..)))............ ( -27.70) >DroEre_CAF1 2197 109 - 1 GGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCAGCAACACACUGCCA-ACUC---GCAACAACUUCUCAGUUCCUCAUCUGCAACUGC-------AACUGCAUCUUUGGCUCC ((((....(((((((.((.(.((((......))))).)).)))))))..)))).-....---(((..((((....)))).......(((....))-------)..)))............ ( -27.90) >DroYak_CAF1 6277 120 - 1 GGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCAGCAACACACUGCCACACUCGCCGCAACAACUUCUCAGUUCCUCAUCGGCAACUGCAACUUGCAACUGCAUCUUUAGCUCC ((((....(((((((.((.(.((((......))))).)).)))))))..)))).........((..........(((((((.....)).))))).....(((....)))......))... ( -32.60) >consensus GGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA_ACUC___GCAACAACUUCUCAGUUCCUCAUCUGCAACUGC_______AACUGCAUCUUUAGCUCC ((((....(((((((.((.(.((((......))))).)).)))))))..)))).........(((.........(((((.(......).)))))...........)))............ (-28.45 = -28.05 + -0.40)


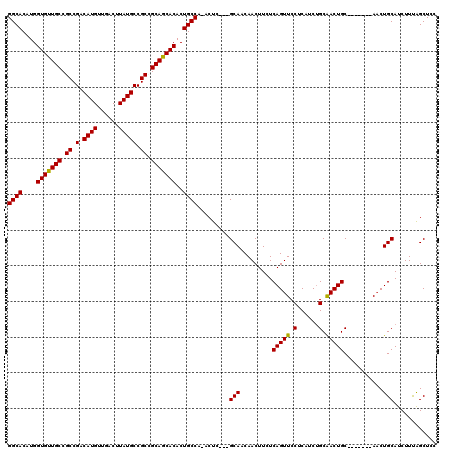
| Location | 14,478,429 – 14,478,524 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.83 |
| Mean single sequence MFE | -36.32 |
| Consensus MFE | -24.80 |
| Energy contribution | -24.91 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.693083 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14478429 95 + 22224390 GAACUGAGAAGUUGUUGC---GAGU-UGGCAGUGUGCUGCGGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG---------------------CAAGAUGCAAUAUG .((((....)))).((((---(...-((((((((((((((..((((((........))))))))))).))))....)))))..))---------------------)))........... ( -31.40) >DroSec_CAF1 3632 95 + 1 GAACUGAGAAGUUGUUGC---GAGU-UGGCAGUGUGCUGCGGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG---------------------CAAGAUGCAAUAUG .((((....)))).((((---(...-((((((((((((((..((((((........))))))))))).))))....)))))..))---------------------)))........... ( -31.40) >DroSim_CAF1 10132 95 + 1 GAACUGAGAAGUUGUUGC---GAGU-UGGCAGUGUGCUGCGGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG---------------------CAAGAUGCAAUAUG .((((....)))).((((---(...-((((((((((((((..((((((........))))))))))).))))....)))))..))---------------------)))........... ( -31.40) >DroEre_CAF1 2230 102 + 1 GAACUGAGAAGUUGUUGC---GAGU-UGGCAGUGUGUUGCUGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUGCAA--------------GCUGCAAGCUGCAAGAUG .((((....)))).((((---.(((-(.((((((((((((((((((((........))))).))))))))))..((..(...)..)).--------------))))).)))))))).... ( -39.90) >DroYak_CAF1 6317 120 + 1 GAACUGAGAAGUUGUUGCGGCGAGUGUGGCAGUGUGUUGCUGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUGCCAUGUGCAAUGGGAGAUGUGCAAGAUGCAAUAUC .((((....))))(((((((((...(((((((((((((((((((((((........))))).))))))))).))).)))))).)))).((..((........))..))....)))))... ( -47.50) >consensus GAACUGAGAAGUUGUUGC___GAGU_UGGCAGUGUGCUGCGGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG_____________________CAAGAUGCAAUAUG .((((....))))(((((........((((((((((((((..((((((........))))))))))).))))....)))))..(((...................)))....)))))... (-24.80 = -24.91 + 0.11)

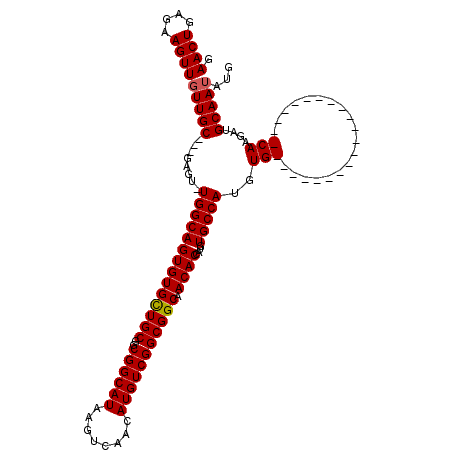
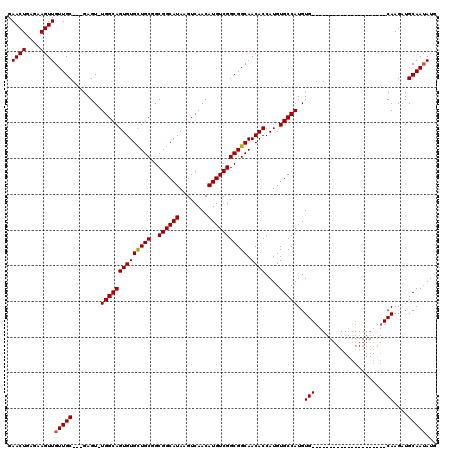
| Location | 14,478,429 – 14,478,524 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.83 |
| Mean single sequence MFE | -28.44 |
| Consensus MFE | -24.22 |
| Energy contribution | -24.13 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.68 |
| SVM RNA-class probability | 0.971579 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14478429 95 - 22224390 CAUAUUGCAUCUUG---------------------CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA-ACUC---GCAACAACUUCUCAGUUC ...........(((---------------------(...(((((....(((((((.((.(.((((......))))).)).)))))))..)))))-....---)))).............. ( -26.40) >DroSec_CAF1 3632 95 - 1 CAUAUUGCAUCUUG---------------------CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA-ACUC---GCAACAACUUCUCAGUUC ...........(((---------------------(...(((((....(((((((.((.(.((((......))))).)).)))))))..)))))-....---)))).............. ( -26.40) >DroSim_CAF1 10132 95 - 1 CAUAUUGCAUCUUG---------------------CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA-ACUC---GCAACAACUUCUCAGUUC ...........(((---------------------(...(((((....(((((((.((.(.((((......))))).)).)))))))..)))))-....---)))).............. ( -26.40) >DroEre_CAF1 2230 102 - 1 CAUCUUGCAGCUUGCAGC--------------UUGCACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCAGCAACACACUGCCA-ACUC---GCAACAACUUCUCAGUUC ....((((....(((...--------------..)))..(((((....(((((((.((.(.((((......))))).)).)))))))..)))))-....---)))).............. ( -29.70) >DroYak_CAF1 6317 120 - 1 GAUAUUGCAUCUUGCACAUCUCCCAUUGCACAUGGCACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCAGCAACACACUGCCACACUCGCCGCAACAACUUCUCAGUUC ....((((....((((..........))))...(((...(((((....(((((((.((.(.((((......))))).)).)))))))..))))).....))))))).............. ( -33.30) >consensus CAUAUUGCAUCUUG_____________________CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCCGCAGCACACUGCCA_ACUC___GCAACAACUUCUCAGUUC ....((((....(((...................)))..(((((....(((((((.((.(.((((......))))).)).)))))))..)))))........)))).............. (-24.22 = -24.13 + -0.09)

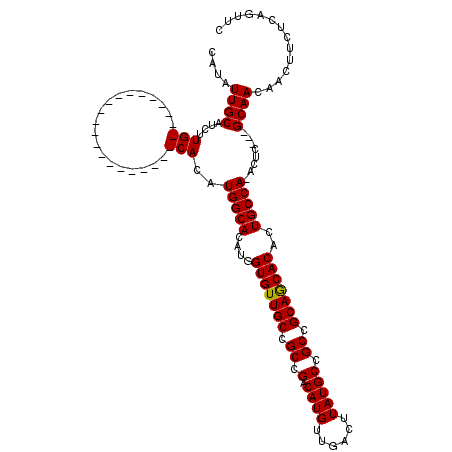
| Location | 14,478,465 – 14,478,562 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 85.88 |
| Mean single sequence MFE | -33.77 |
| Consensus MFE | -22.41 |
| Energy contribution | -24.10 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.863865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14478465 97 + 22224390 GGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG-------CAAGAUGCAAUAUGCAAUGUUGCUGCUUCUGCUGC--CUGCUGCCUGCAUAAUU (((((((.((((((((((...((.(....).))...(((....((-------(.....)))....)))))))))..)))).))))))--)(((.....)))..... ( -34.60) >DroSec_CAF1 3668 90 + 1 GGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG-------CAAGAUGCAAUAUGCAAUGUUGCUGCUUCUGCU---------GCCUGCAUAAUU (((((((.((((((((((...((.(....).))...(((....((-------(.....)))....)))))))))..)))).))))---------)))......... ( -32.50) >DroSim_CAF1 10168 97 + 1 GGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG-------CAAGAUGCAAUAUGCAAUGUUGCUGCUUCUGCUGC--CUGCUGCCUGCAUAAUU (((((((.((((((((((...((.(....).))...(((....((-------(.....)))....)))))))))..)))).))))))--)(((.....)))..... ( -34.60) >DroEre_CAF1 2266 97 + 1 UGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUGCAAGCUGCAAGCUGCAAGAUGCAAUGUUGC---------UGCUGCUACUGCCUGCAUAAUU (((((((...(....)..(.(((((((((((.....(((....((((.((.....))))))....))).))))))---------))))))...)))).)))..... ( -33.40) >consensus GGCGGCAUAAGUCAACAUGUCGGCGGCAACACCAUGUGCCAUGUG_______CAAGAUGCAAUAUGCAAUGUUGCUGCUUCUGCUGC__CUGCUGCCUGCAUAAUU ((((((((........)))))((((((((((.(((((((....(((.....)))....)).)))))...))))))))))...............)))......... (-22.41 = -24.10 + 1.69)


| Location | 14,478,465 – 14,478,562 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 85.88 |
| Mean single sequence MFE | -30.62 |
| Consensus MFE | -21.48 |
| Energy contribution | -23.22 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.611358 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14478465 97 - 22224390 AAUUAUGCAGGCAGCAG--GCAGCAGAAGCAGCAACAUUGCAUAUUGCAUCUUG-------CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCC ......((.((((..((--((((((...((.((((((((((....(((.....)-------))....)))....))))))).)).....))))).))).)))))). ( -32.00) >DroSec_CAF1 3668 90 - 1 AAUUAUGCAGGC---------AGCAGAAGCAGCAACAUUGCAUAUUGCAUCUUG-------CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCC .........(((---------.(((.(((((((..((((((....(((.....)-------))....))).)))((((((....)))))))))).))).))).))) ( -26.70) >DroSim_CAF1 10168 97 - 1 AAUUAUGCAGGCAGCAG--GCAGCAGAAGCAGCAACAUUGCAUAUUGCAUCUUG-------CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCC ......((.((((..((--((((((...((.((((((((((....(((.....)-------))....)))....))))))).)).....))))).))).)))))). ( -32.00) >DroEre_CAF1 2266 97 - 1 AAUUAUGCAGGCAGUAGCAGCA---------GCAACAUUGCAUCUUGCAGCUUGCAGCUUGCACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCA .....(((.((((.(((((((.---------(((((((((((..((((.....))))..))))((((...))))))))))).)).....))))).....))))))) ( -31.80) >consensus AAUUAUGCAGGCAGCAG__GCAGCAGAAGCAGCAACAUUGCAUAUUGCAUCUUG_______CACAUGGCACAUGGUGUUGCCGCCGACAUGUUGACUUAUGCCGCC ......((.((((.......(((((...((.((((((((((.....)))..............((((...))))))))))).)).....))))).....)))))). (-21.48 = -23.22 + 1.75)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:52:24 2006