
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,412,992 – 14,413,176 |
| Length | 184 |
| Max. P | 0.977612 |

| Location | 14,412,992 – 14,413,112 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.92 |
| Mean single sequence MFE | -34.04 |
| Consensus MFE | -28.69 |
| Energy contribution | -29.37 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.39 |
| SVM RNA-class probability | 0.948917 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14412992 120 + 22224390 AAUUACGUAUGCAGAAAAUGACCAACAUGUUGCUCCCAGGACUGUGGCAUAAAAAAUUGGCAUAGGAAUGAGGCAGUUGCAAGGGGCAAGGGCAGGAUGUCAUAUAGAAAUGGCAGCAUG .......(((((.........((....(.(((((((((...(((((.((........)).)))))...))..((....))..))))))).)...)).((((((......))))))))))) ( -32.90) >DroSec_CAF1 122712 120 + 1 AAUUACGUAUGCAGAAAAUGACCAACAUGUUGCUCCCAGGACUGUGCCAUAAAAAAUUGGCAUAAGAAUGGGGCAGAUGCAAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUG .......(((((......((.((......((((.((((....(((((((........)))))))....)))))))).(((.....))).)).))...((((((......))))))))))) ( -38.10) >DroSim_CAF1 115909 120 + 1 AAUUACGUAUGCAGAAAAUGACCAACAUGUUGCUCCCAGGACUGUGCCAUAAAAAAUUGGCAUAAGAAUGAGGCAGAUGCAAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUG .......(((((.........((....(.(((((((((....(((((((........)))))))....))..((....))..))))))).)...)).((((((......))))))))))) ( -36.00) >DroEre_CAF1 107732 117 + 1 AAUUACGUAUGCAGAAAAUGACCAACAUGUUGCUCCCAGGACUCUGCCAUAAAAAAUUGGCAC-AGAAUAAGGCAGAAGU--GGGGCGAGGGCACGAUGCCAUGUAGAAAUUGCAGCAUG .........................((((((((.((((....((((((...............-.......))))))..)--)))(((..(((.....))).))).......)))))))) ( -29.15) >consensus AAUUACGUAUGCAGAAAAUGACCAACAUGUUGCUCCCAGGACUGUGCCAUAAAAAAUUGGCAUAAGAAUGAGGCAGAUGCAAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUG .......(((((.........((....(.((((.(((.....(((((((........)))))))........((....))..))))))).)...)).((((((......))))))))))) (-28.69 = -29.37 + 0.69)


| Location | 14,413,032 – 14,413,152 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.98 |
| Mean single sequence MFE | -32.42 |
| Consensus MFE | -26.41 |
| Energy contribution | -26.72 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.80 |
| SVM RNA-class probability | 0.977612 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
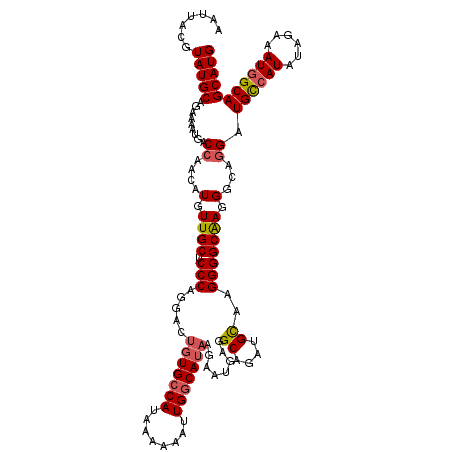
>X_DroMel_CAF1 14413032 120 + 22224390 ACUGUGGCAUAAAAAAUUGGCAUAGGAAUGAGGCAGUUGCAAGGGGCAAGGGCAGGAUGUCAUAUAGAAAUGGCAGCAUGCAUACUAAAAAUGCACUUCUGAGAUGCACUGAAAGCAAAC .(((((.((........)).)))))...(.(((((....((..(((((((.(((...((((((......))))))...)))...)).....))).))..))...))).)).)........ ( -29.70) >DroSec_CAF1 122752 119 + 1 ACUGUGCCAUAAAAAAUUGGCAUAAGAAUGGGGCAGAUGCAAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUGCAUACUAA-AAUGCACUUCUGAGAUGCACUGAAAGCAAAA ..(((((((........)))))))....(.(((((....((..(((((((.(((...((((((......))))))...)))...))..-..))).))..))...))).)).)........ ( -33.90) >DroSim_CAF1 115949 119 + 1 ACUGUGCCAUAAAAAAUUGGCAUAAGAAUGAGGCAGAUGCAAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUGCAUACUAA-AAUGCACUUCUGAGAUGCACUGAAAGCAAAA ..(((((((........)))))))....(.(((((....((..(((((((.(((...((((((......))))))...)))...))..-..))).))..))...))).)).)........ ( -33.90) >DroEre_CAF1 107772 115 + 1 ACUCUGCCAUAAAAAAUUGGCAC-AGAAUAAGGCAGAAGU--GGGGCGAGGGCACGAUGCCAUGUAGAAAUUGCAGCAUGCAUACUAA-GCUGCACUUCUAAGAUGCACUAAA-GAAGAC .((.(((((........))))).-))....((((((((((--(.(((...(((.....)))...(((....(((.....)))..))).-))).)))))).....))).))...-...... ( -32.20) >consensus ACUGUGCCAUAAAAAAUUGGCAUAAGAAUGAGGCAGAUGCAAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUGCAUACUAA_AAUGCACUUCUGAGAUGCACUGAAAGCAAAA ..(((((((........)))))))......(((((....((..(((((((.(((...((((((......))))))...)))...)).....))).))..))...))).)).......... (-26.41 = -26.72 + 0.31)



| Location | 14,413,072 – 14,413,176 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 77.38 |
| Mean single sequence MFE | -26.92 |
| Consensus MFE | -15.29 |
| Energy contribution | -15.60 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792834 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
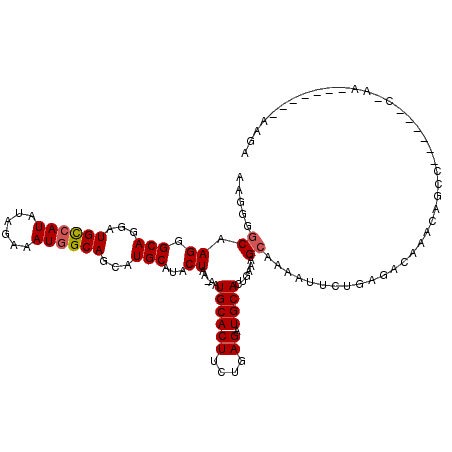
>X_DroMel_CAF1 14413072 104 + 22224390 AAGGGGCAAGGGCAGGAUGUCAUAUAGAAAUGGCAGCAUGCAUACUAAAAAUGCACUUCUGAGAUGCACUGAAAGCAAACUUCUGAGACAAAUAUCU------C-AA-------UCGA (((..((.((.(((...((((((......))))))...)))...)).....((((((....)).))))......))...))).(((((......)))------)-).-------.... ( -21.40) >DroSec_CAF1 122792 103 + 1 AAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUGCAUACUAA-AAUGCACUUCUGAGAUGCACUGAAAGCAAAAUUCUGAGAAAAACAGCC------C-AA-------AAGA ...((((.((.(((...((((((......))))))...)))...))..-.......((((.((((((.......)))....))).)))).....)))------)-..-------.... ( -29.70) >DroSim_CAF1 115989 103 + 1 AAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUGCAUACUAA-AAUGCACUUCUGAGAUGCACUGAAAGCAAAAAUCUGAGACAAACAGCC------C-AA-------AAGC ...((((.((.(((...((((((......))))))...)))...))..-........(((.((((((.......)))....))).)))......)))------)-..-------.... ( -28.60) >DroEre_CAF1 107811 109 + 1 --GGGGCGAGGGCACGAUGCCAUGUAGAAAUUGCAGCAUGCAUACUAA-GCUGCACUUCUAAGAUGCACUAAA-GAAGACCUC-----CGAUUGUCUGUGGCCUUAAAUAAGCACAGC --.....((((.(((..(((....(((((..((((((...........-)))))).)))))....))).....-..((((...-----.....))))))).))))............. ( -28.00) >consensus AAGGGGCAAGGGCAGGAUGCCAUAUAGAAAUGGCAGCAUGCAUACUAA_AAUGCACUUCUGAGAUGCACUGAAAGCAAAAUUCUGAGACAAACAGCC______C_AA_______AAGA .....((.((.(((...((((((......))))))...)))...)).....((((((....)).))))......)).......................................... (-15.29 = -15.60 + 0.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:51:21 2006