
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,380,603 – 14,380,750 |
| Length | 147 |
| Max. P | 0.990312 |

| Location | 14,380,603 – 14,380,711 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 86.92 |
| Mean single sequence MFE | -25.50 |
| Consensus MFE | -19.12 |
| Energy contribution | -20.12 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.778936 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14380603 108 + 22224390 --CACAUGCACGCAUAUUCUCGUCCUGCGACGAUGCAUAUAUGUAUGUGAAUGGAUCUGUGUGAGUGAGUUGGCUCACGUCAGCUAGACAUAAACCAAACAAUAAACAAA --((((((((...((((.((((((....))))).).)))).))))))))..(((......((((((......))))))(((.....))).....)))............. ( -28.90) >DroSec_CAF1 96632 104 + 1 --CACACACACGCACAUUUUAGUCCUGCGACGAUGCAUAUA----UGUGAAUGGGUCUGUGUGAGUGAGUUGGCUCACGUCAGCUAGACAUAAACCAAACAAUAAACAAA --....(((.((((((.....(((....)))(((.(((...----.....))).))))))))).)))(((((((....)))))))......................... ( -25.30) >DroSim_CAF1 90461 104 + 1 --CACACACACGCACAUUCUAGUCCUGCGACGAUGCAUAUA----UGUGAAUGGGUCUGUGUGAGUGAGUUGGCUCACGUCAGCUAGACAUAAACCAAACAAUAAACAAA --..((((((.((.(((((..(((....)))...(((....----)))))))).)).)))))).(((((....)))))(((.....)))..................... ( -27.10) >DroYak_CAF1 98580 94 + 1 CACACACGCACACACAUUCUAGUCCUGCGACCAUGUAUACA----U------------GUGUGAGUGAGUUGGCUCACGUCAGCUAGACAUAAACCAAACAAUAAACAAA (((((.((((...((......))..))))...(((....))----)------------))))).(((((....)))))(((.....)))..................... ( -20.70) >consensus __CACACACACGCACAUUCUAGUCCUGCGACGAUGCAUAUA____UGUGAAUGGGUCUGUGUGAGUGAGUUGGCUCACGUCAGCUAGACAUAAACCAAACAAUAAACAAA ....(((.(((((........(((....)))(((.(((............))).))).))))).)))(((((((....)))))))......................... (-19.12 = -20.12 + 1.00)


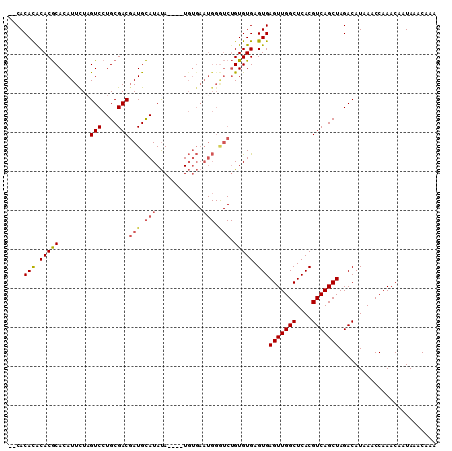
| Location | 14,380,603 – 14,380,711 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 86.92 |
| Mean single sequence MFE | -34.62 |
| Consensus MFE | -26.79 |
| Energy contribution | -26.97 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.13 |
| Mean z-score | -3.45 |
| Structure conservation index | 0.77 |
| SVM decision value | 2.21 |
| SVM RNA-class probability | 0.990312 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14380603 108 - 22224390 UUUGUUUAUUGUUUGGUUUAUGUCUAGCUGACGUGAGCCAACUCACUCACACAGAUCCAUUCACAUACAUAUAUGCAUCGUCGCAGGACGAGAAUAUGCGUGCAUGUG-- (((((....((.((((((((((((.....))))))))))))..)).....)))))......(((((.((..((((..(((((....)))))...))))..)).)))))-- ( -31.90) >DroSec_CAF1 96632 104 - 1 UUUGUUUAUUGUUUGGUUUAUGUCUAGCUGACGUGAGCCAACUCACUCACACAGACCCAUUCACA----UAUAUGCAUCGUCGCAGGACUAAAAUGUGCGUGUGUGUG-- (((((....((.((((((((((((.....))))))))))))..)).....)))))......((((----(((((((((.(((....)))......)))))))))))))-- ( -33.60) >DroSim_CAF1 90461 104 - 1 UUUGUUUAUUGUUUGGUUUAUGUCUAGCUGACGUGAGCCAACUCACUCACACAGACCCAUUCACA----UAUAUGCAUCGUCGCAGGACUAGAAUGUGCGUGUGUGUG-- (((((....((.((((((((((((.....))))))))))))..)).....)))))......((((----(((((((((..((.........))..)))))))))))))-- ( -34.40) >DroYak_CAF1 98580 94 - 1 UUUGUUUAUUGUUUGGUUUAUGUCUAGCUGACGUGAGCCAACUCACUCACAC------------A----UGUAUACAUGGUCGCAGGACUAGAAUGUGUGUGCGUGUGUG ............((((((((((((.....))))))))))))......(((((------------(----(((((((((((((....)))).....))))))))))))))) ( -38.60) >consensus UUUGUUUAUUGUUUGGUUUAUGUCUAGCUGACGUGAGCCAACUCACUCACACAGACCCAUUCACA____UAUAUGCAUCGUCGCAGGACUAGAAUGUGCGUGCGUGUG__ ............((((((((((((.....))))))))))))......(((((.((.....))........((((((((((((....)))).....))))))))))))).. (-26.79 = -26.97 + 0.19)

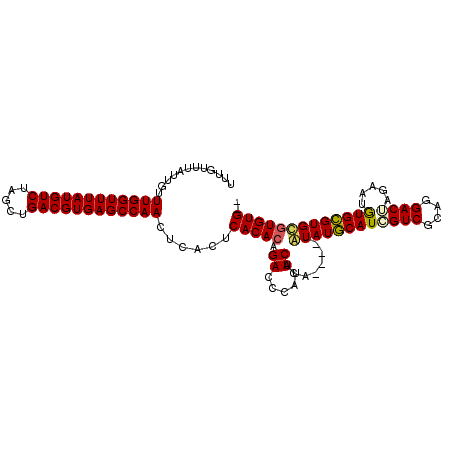
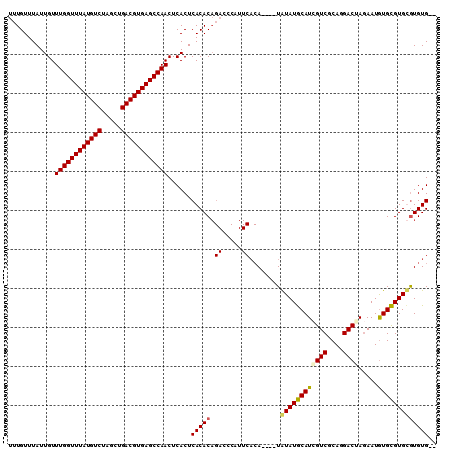
| Location | 14,380,640 – 14,380,750 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 87.75 |
| Mean single sequence MFE | -22.52 |
| Consensus MFE | -19.01 |
| Energy contribution | -19.20 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.825075 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14380640 110 - 22224390 UACUUGCCAGCAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUCUAGCU-GACGUGAGCCAACU-CACUCACACAGAUCCAUUCACAUACAUA .......((((....(((..(((((((((((.((((((....))))))))))))))))))))...)))-)..(((((......-..)))))..................... ( -21.70) >DroSec_CAF1 96669 106 - 1 UACUUGCCAGCAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUCUAGCU-GACGUGAGCCAACU-CACUCACACAGACCCAUUCACA----UA .......((((....(((..(((((((((((.((((((....))))))))))))))))))))...)))-)..(((((......-..)))))...............----.. ( -21.70) >DroSim_CAF1 90498 106 - 1 UACUUGCCAGCAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUCUAGCU-GACGUGAGCCAACU-CACUCACACAGACCCAUUCACA----UA .......((((....(((..(((((((((((.((((((....))))))))))))))))))))...)))-)..(((((......-..)))))...............----.. ( -21.70) >DroEre_CAF1 86386 106 - 1 UACUUGCCGGCAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUAUAGCU-GACGUGAGCCAACU-CACGCACACAGACGCACUCGCA----UG ....(((((((....(((..(((((((((((.((((((....))))))))))))))))))))...)))-).((((((....))-))))...............)))----.. ( -24.60) >DroYak_CAF1 98619 94 - 1 UACUUGCCAGCAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUCUAGCU-GACGUGAGCCAACU-CACUCACAC------------A----UG .......((((....(((..(((((((((((.((((((....))))))))))))))))))))...)))-)..(((((......-..)))))..------------.----.. ( -21.70) >DroAna_CAF1 163507 96 - 1 UACUUGCCAACAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUCUAGCUAGACGUGGGCCAAGUACACUCAUCC------------G----AG ((((((((.((.........(((((((((((.((((((....))))))))))))))))).((((....)))))).)).))))))..(((....------------)----)) ( -23.70) >consensus UACUUGCCAGCAUUCACAACUAAAUUAGAUGCAUAAACUUUUGUUUAUUGUUUGGUUUAUGUCUAGCU_GACGUGAGCCAACU_CACUCACACAGACCCAUUCACA____UA .........(((((.............)))))((((((....))))))...((((((((((((......))))))))))))............................... (-19.01 = -19.20 + 0.20)


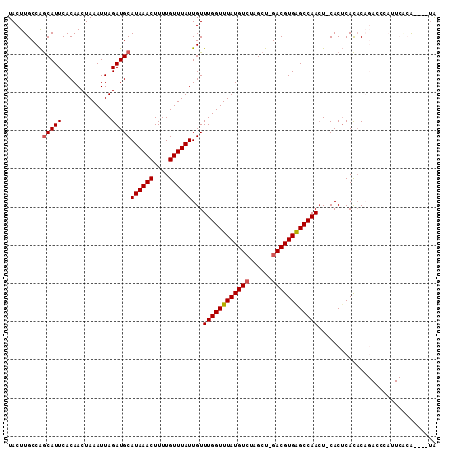
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:59 2006