| Sequence ID | X_DroMel_CAF1 |
|---|---|
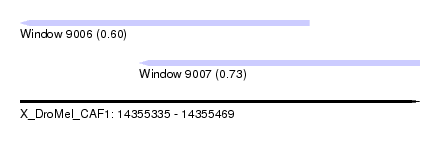
| Location | 14,355,335 – 14,355,469 |
| Length | 134 |
| Max. P | 0.728585 |

| Location | 14,355,335 – 14,355,432 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 87.54 |
| Mean single sequence MFE | -22.05 |
| Consensus MFE | -13.99 |
| Energy contribution | -14.42 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.598410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14355335 97 - 22224390 -AAAAUGGCAUAUUUGCCA-----------AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUAUUGAGUUAGUUUACCAUGUAAUUAGGCAACCUGUUGUACA -....(((((....)))))-----------...((((((((.((((((((....))))))))((((.....))))..))))))))((((..((((...))))...)))) ( -22.60) >DroSec_CAF1 69837 97 - 1 -AAAAUGACAUAUUUGCCA-----------AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUAUUUAGUUAGUUUACCAUGUAAUUAGGCAACCUCUUGUACA -......(((...(((((.-----------.((((((((((.((((((((....)))))))).....((......)))))))))).)).....))))).....)))... ( -17.50) >DroSim_CAF1 65117 97 - 1 -AAAAUGACAUAUUUGCCA-----------AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUAUUGAGUUAGUUUACCAUGUAAUUAGGCAACCUGUUGUGCA -...(..(((...(((((.-----------.((((((((((.((((((((....))))))))((((.....))))..)))))))).)).....)))))..)))..)... ( -20.90) >DroYak_CAF1 72017 109 - 1 AAAAAUGGCACAUUCGCCAUCGUUUUGCCAAGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGCAUUCAGUUUGUUUACCUUGUAAUUAGGCAACCCAGUGUAAA ....(((((......))))).(..((((((((((((((((((.(((((((....))))))))))))))...))))))..((((...))))...)))))..)........ ( -27.20) >consensus _AAAAUGACAUAUUUGCCA___________AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUAUUGAGUUAGUUUACCAUGUAAUUAGGCAACCUGUUGUACA .......(((...(((((............((((((((((((.(((((((....)))))))))))))...))))))...((((...))))...))))).....)))... (-13.99 = -14.42 + 0.44)



| Location | 14,355,375 – 14,355,469 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 86.70 |
| Mean single sequence MFE | -20.85 |
| Consensus MFE | -13.93 |
| Energy contribution | -13.99 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.728585 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14355375 94 - 22224390 GUGCUUAACCGACCCUAAUGUUUAUGCAGUAUUUACA----AAAAUGGCAUAUUUGCCA-----------AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUA ..........(((.....(((((((.((((((((((.----....(((((....)))))-----------.....)))))).)))))))))))...))).......... ( -20.10) >DroSec_CAF1 69877 94 - 1 GUGCUUAACUGUCCCUAAUGUUUAUGCAGUAUUUACA----AAAAUGACAUAUUUGCCA-----------AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUA .(((..(((..........)))...))).....((((----((.......((((((((.-----------....))))))))((((((((....)))))))).)))))) ( -18.90) >DroSim_CAF1 65157 94 - 1 GUGCUUAACCGUCCCUAAUGUUUAUGCAGUAUUUACA----AAAAUGACAUAUUUGCCA-----------AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUA .(((.(((.(((.....))).))).))).....((((----((.......((((((((.-----------....))))))))((((((((....)))))))).)))))) ( -19.30) >DroYak_CAF1 72057 109 - 1 GUGCUCAACCGCCCCGGAUGUUUAUGCAAUAAUUACACAAAAAAAUGGCACAUUCGCCAUCGUUUUGCCAAGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGCA .(((.((.(((...))).)).....))).........(((((..(((((......)))))..)))))........(((((((.(((((((....)))))))))))))). ( -25.10) >consensus GUGCUUAACCGUCCCUAAUGUUUAUGCAGUAUUUACA____AAAAUGACAUAUUUGCCA___________AGCUGGUAAAUAAUUGAUAAAUAAUUGUCAAUAUUUGUA .(((..(((..........)))...)))...................((((((((((((..............)))))))))((((((((....))))))))...))). (-13.93 = -13.99 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:41 2006