
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,310,787 – 14,310,883 |
| Length | 96 |
| Max. P | 0.875689 |

| Location | 14,310,787 – 14,310,883 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 93.09 |
| Mean single sequence MFE | -31.56 |
| Consensus MFE | -26.06 |
| Energy contribution | -26.30 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.658477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14310787 96 + 22224390 AUUUU------UUGGCCAACGUCGGAGGCGCUAAUUAAUCCAUUAAGGCCAAGUUUGUUGCGUGGUGUAAUUGGCAUUGCGGUUGCGCAGAAAACGUUGGCG .....------...((((((((.....((((....(((((((...(.((((.((.....)).)))).)...))).)))).....)))).....)))))))). ( -31.60) >DroSec_CAF1 23535 96 + 1 AUUUU------UUGGCCAACGGCGGAGGCGCUAAUUAAUCCAUUAAGGCCAAGUUUGUUGCGCGGUGUAAUUGUCAUUGCGGUUGCCCAGAAAACGUUGGCG .....------...(((((((......((((.((((...((.....))...))))....))))((.((((((((....))))))))))......))))))). ( -30.30) >DroSim_CAF1 20840 96 + 1 AUUUU------UUGGCCAACGGCGGAGGCGCUAAUUAAUCCAUUAAGGCCAAGUUUGUUGCGCGGUGUAAUUGGCAUUGCGGUUGCCCAGAAAACGUUGGCG .....------...(((((((......((((.((((...((.....))...))))....))))((.(((((((......)))))))))......))))))). ( -29.40) >DroEre_CAF1 20444 96 + 1 GCUUU------UCGGCCAACGGAGGAGGCGCUAAUUAGUCCAUUAAGGCCAAGUUUGUUGCGCGGUGUAAUUGGCAUUGCGGUUGCCCAGAAAACGUUGGCG .....------...(((((((......((((.((((.(.((.....)).).))))....))))((.(((((((......)))))))))......))))))). ( -30.60) >DroYak_CAF1 24800 102 + 1 GUUUUUUGUUUUUGGGCAACGGAGGAGGCGCUAAUUAAUCCAUUAAGGCCAAGUUUGUUGCGCGGUGUAAUUGGCAUUGCGGUUGCCCAGAAAACGUUGGCG (((.....((((((((((((......(((..((((......))))..)))..........((((((((.....)))))))))))))))))))).....))). ( -35.90) >consensus AUUUU______UUGGCCAACGGCGGAGGCGCUAAUUAAUCCAUUAAGGCCAAGUUUGUUGCGCGGUGUAAUUGGCAUUGCGGUUGCCCAGAAAACGUUGGCG ..............(((((((......((((.((((...((.....))...))))....))))((.(((((((......)))))))))......))))))). (-26.06 = -26.30 + 0.24)

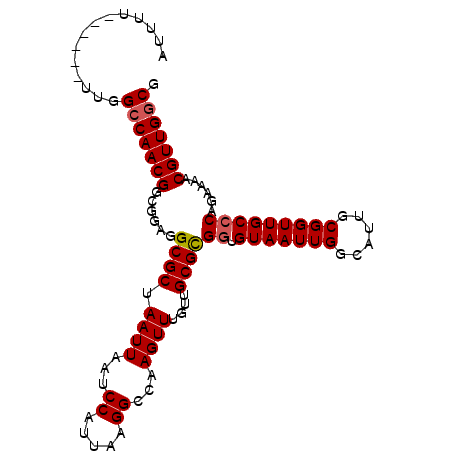
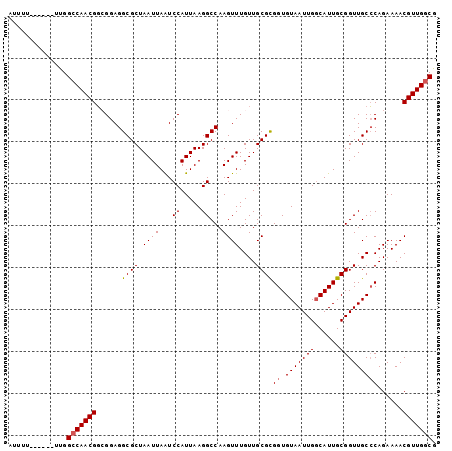
| Location | 14,310,787 – 14,310,883 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 93.09 |
| Mean single sequence MFE | -25.25 |
| Consensus MFE | -19.36 |
| Energy contribution | -19.76 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.875689 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14310787 96 - 22224390 CGCCAACGUUUUCUGCGCAACCGCAAUGCCAAUUACACCACGCAACAAACUUGGCCUUAAUGGAUUAAUUAGCGCCUCCGACGUUGGCCAA------AAAAU .(((((((((....((((.........(((((..................)))))..((((......))))))))....)))))))))...------..... ( -25.87) >DroSec_CAF1 23535 96 - 1 CGCCAACGUUUUCUGGGCAACCGCAAUGACAAUUACACCGCGCAACAAACUUGGCCUUAAUGGAUUAAUUAGCGCCUCCGCCGUUGGCCAA------AAAAU .(((((((((...(((....)))....))).........(((..........(((.(((((......))))).)))..))).))))))...------..... ( -23.90) >DroSim_CAF1 20840 96 - 1 CGCCAACGUUUUCUGGGCAACCGCAAUGCCAAUUACACCGCGCAACAAACUUGGCCUUAAUGGAUUAAUUAGCGCCUCCGCCGUUGGCCAA------AAAAU .(((((((......((....))((..((((.........).)))........(((.(((((......))))).)))...)))))))))...------..... ( -25.30) >DroEre_CAF1 20444 96 - 1 CGCCAACGUUUUCUGGGCAACCGCAAUGCCAAUUACACCGCGCAACAAACUUGGCCUUAAUGGACUAAUUAGCGCCUCCUCCGUUGGCCGA------AAAGC .(((((((.......((((.......)))).........((((.......((((((.....)).))))...))))......)))))))...------..... ( -25.00) >DroYak_CAF1 24800 102 - 1 CGCCAACGUUUUCUGGGCAACCGCAAUGCCAAUUACACCGCGCAACAAACUUGGCCUUAAUGGAUUAAUUAGCGCCUCCUCCGUUGCCCAAAAACAAAAAAC .......(((((.(((((((((((..((.......))..)))..........(((.(((((......))))).)))......)))))))))))))....... ( -26.20) >consensus CGCCAACGUUUUCUGGGCAACCGCAAUGCCAAUUACACCGCGCAACAAACUUGGCCUUAAUGGAUUAAUUAGCGCCUCCGCCGUUGGCCAA______AAAAU .(((((((......((....))..............................(((.(((((......))))).))).....))))))).............. (-19.36 = -19.76 + 0.40)

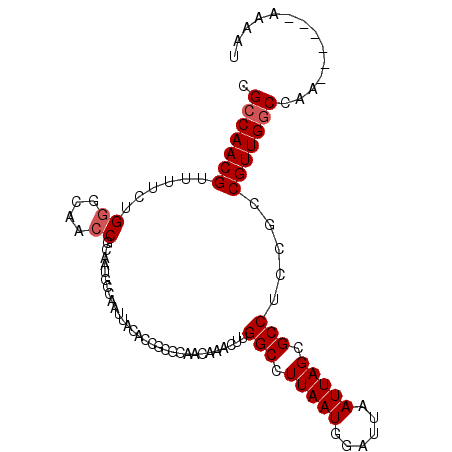

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:17 2006