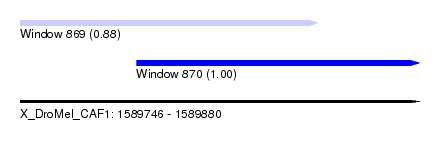
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,589,746 – 1,589,880 |
| Length | 134 |
| Max. P | 0.996617 |

| Location | 1,589,746 – 1,589,846 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 82.51 |
| Mean single sequence MFE | -38.83 |
| Consensus MFE | -30.61 |
| Energy contribution | -30.73 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.881250 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
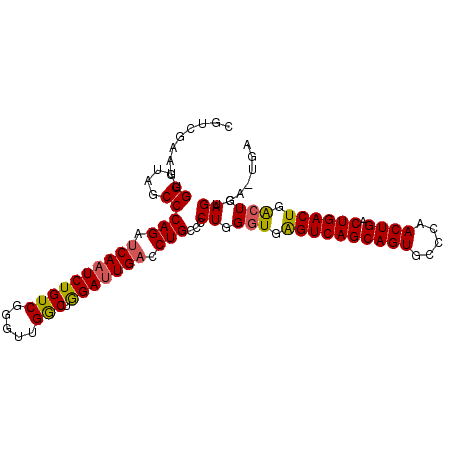
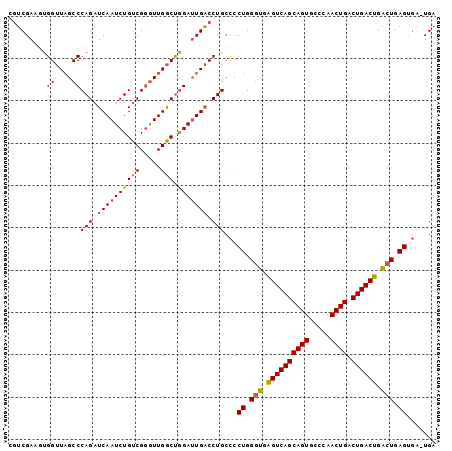
>X_DroMel_CAF1 1589746 100 + 22224390 -GUCGAAAUGGUUAGCCCAGAUCAAUCUGUCGGGUUGGCUAGAUUGACCUGCCUCUGGGUGGGUCAGCAGUGCCCAACUGACUGACUGACUGAGUGAAUGA -(((((..((((((((((.(((......)))))))))))))..)))))....(.((.(((.((((((((((.....)))).)))))).))).)).)..... ( -42.00) >DroSec_CAF1 51193 100 + 1 CGUCGAAGUGGUU-GCCCAGAUCAAUCUGUCGGGUUGGCUGGAGUGACCUGCCCCUGGGUGAGUCAGCAGUGCCCAACUGACUGACUGACUGAGUGAGUGA .........(((.-.(((((.((((((.....)))))))))).)..))).((.(((.(((.((((((((((.....)))).)))))).))).)).).)).. ( -41.90) >DroYak_CAF1 51134 97 + 1 CUGCGAUGUGGCUAGCCCAGCUCAAUCUGUCGCAUUGACCGGAUUGUCCUGCUCCUGGCUGAGUCAGCAGUGUCCAACUGACUGACUGGAUGAG----UGA ..((.....((((((..(((..(((((((((.....)).)))))))..)))...)))))).((((((((((.....)))).))))))......)----).. ( -32.60) >consensus CGUCGAAGUGGUUAGCCCAGAUCAAUCUGUCGGGUUGGCUGGAUUGACCUGCCCCUGGGUGAGUCAGCAGUGCCCAACUGACUGACUGACUGAGUGA_UGA .........((....))(((.((((((((((.....))).))))))).)))...((.(((.((((((((((.....)))).)))))).))).))....... (-30.61 = -30.73 + 0.12)

| Location | 1,589,785 – 1,589,880 |
|---|---|
| Length | 95 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 75.17 |
| Mean single sequence MFE | -31.97 |
| Consensus MFE | -22.94 |
| Energy contribution | -24.00 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.72 |
| SVM decision value | 2.72 |
| SVM RNA-class probability | 0.996617 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
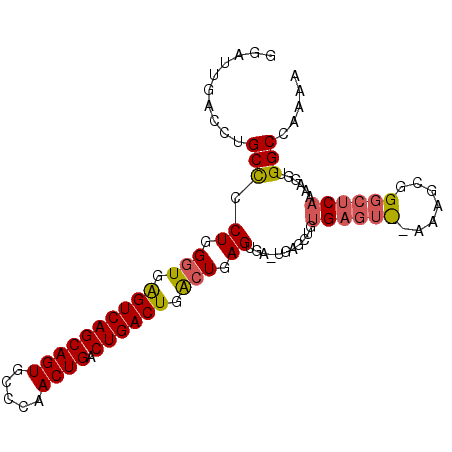
>X_DroMel_CAF1 1589785 95 + 22224390 AGAUUGACCUGCCUCUGGGUGGGUCAGCAGUGCCCAACUGACUGACUGACUGAGUGAAUGAG---UGAGUUGAAAGCGGGCUCAAAAGUUGGCCAAAA ...(((.((.(((.((.(((.((((((((((.....)))).)))))).))).)).)..((((---(..((.....))..)))))...)).)).))).. ( -32.70) >DroSec_CAF1 51232 98 + 1 GGAGUGACCUGCCCCUGGGUGAGUCAGCAGUGCCCAACUGACUGACUGACUGAGUGAGUGAGCUGUGAGUCAAAAGCGGGCUCAAAAGGUGGCCAAAA .(.((.((((....((.(((.((((((((((.....)))).)))))).))).))(((((..(((..........)))..)))))..)))).))).... ( -38.00) >DroYak_CAF1 51174 79 + 1 GGAUUGUCCUGCUCCUGGCUGAGUCAGCAGUGUCCAACUGACUGACUGGAUGAG----UGAGCUG--AGU-------------GAAAGGUGGCAAAAA ...(((((((((((((.(.(.((((((((((.....)))).)))))).).).))----.))))..--...-------------....)).)))))... ( -25.20) >consensus GGAUUGACCUGCCCCUGGGUGAGUCAGCAGUGCCCAACUGACUGACUGACUGAGUGA_UGAGCUGUGAGU__AAAGCGGGCUCAAAAGGUGGCCAAAA ..........(((.((.(((.((((((((((.....)))).)))))).))).))...........((((((.......))))))......)))..... (-22.94 = -24.00 + 1.06)

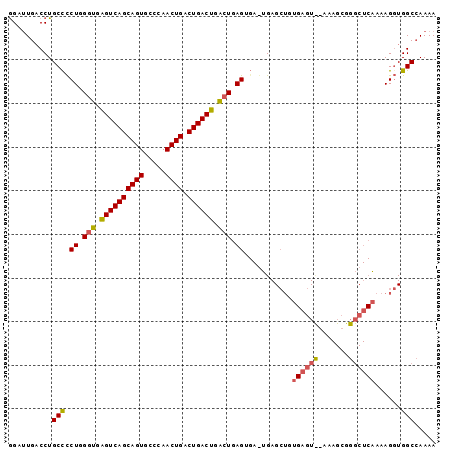
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:47:34 2006