
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,297,310 – 14,297,418 |
| Length | 108 |
| Max. P | 0.930991 |

| Location | 14,297,310 – 14,297,418 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 85.16 |
| Mean single sequence MFE | -29.44 |
| Consensus MFE | -18.44 |
| Energy contribution | -19.44 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.89 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.904691 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14297310 108 + 22224390 UUUGCAUAAUGCACGACACUUUUGAAUCAUGAACGUGGACGUGGACAUGAAAGAAGCCUUAACUCCA-UGACCACGU---CGUCCA----AUCCAAUGCAGCCCGACCAAUACUCC .((((((.......(((..........(((....)))(((((((.((((................))-)).))))))---))))..----.....))))))............... ( -23.73) >DroSec_CAF1 10222 111 + 1 UUUGCAUAAUGCACGACACUUUUGAAUCAUGGACGUGGACGUGGACAUGAAAGAAGCCUUAACUCCA-UGACCGCGUCCACGUCCA----AUCCAAUGCAGCCCGACCAAUCCUCC .((((((......(((.....))).....(((((((((((((((.((((................))-)).)))))))))))))))----.....))))))............... ( -38.09) >DroSim_CAF1 11220 111 + 1 UUUGCAUAAUGCACGACACUUUUGAAUCAUGGACGUGGACGUGGACAUGAAAGAAGCCUUAACUCCA-UGACCGCGUCCACGUCCA----AUCCAAUGCAGCCCGACCAAUCCUCC .((((((......(((.....))).....(((((((((((((((.((((................))-)).)))))))))))))))----.....))))))............... ( -38.09) >DroEre_CAF1 9447 99 + 1 UUUGCAUAAUGCACGACACUUUUGAAUCAUGGAC---GACGUGGACAUGAAAGAAGCCUUAACUCCA-UGACCACGA---CGUCCG----AACCAAUCC------UCCAGAAAUCC ..(((.....)))......(((((......((((---(.(((((.((((................))-)).))))).---)))))(----(........------))))))).... ( -22.59) >DroYak_CAF1 12049 98 + 1 UUUGCAUAAUGCACGACACUUUUGAAUCAUGGAC---GACGUGGACAUGAAAGAAGCCUUAACUCCACUGACCACGU---CGUCCAACCCAACCAAUCC------UC------UCC ..(((.....)))................(((((---(((((((.((((................)).)).))))))---)))))).............------..------... ( -24.69) >consensus UUUGCAUAAUGCACGACACUUUUGAAUCAUGGACGUGGACGUGGACAUGAAAGAAGCCUUAACUCCA_UGACCACGU___CGUCCA____AUCCAAUGCAGCCCGACCAAUACUCC ..(((.....)))................((((((((.((((((.((.(((((....)))...))...)).)))))).)))))))).............................. (-18.44 = -19.44 + 1.00)
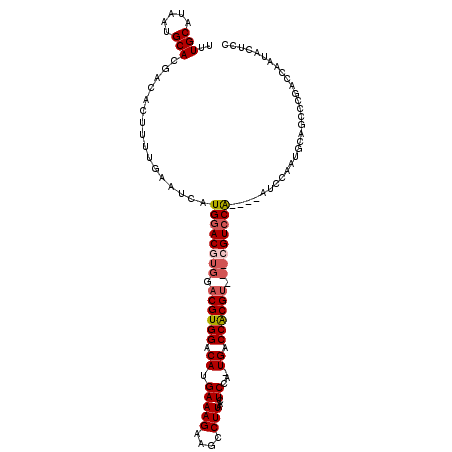


| Location | 14,297,310 – 14,297,418 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 85.16 |
| Mean single sequence MFE | -37.62 |
| Consensus MFE | -26.40 |
| Energy contribution | -28.44 |
| Covariance contribution | 2.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.930991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14297310 108 - 22224390 GGAGUAUUGGUCGGGCUGCAUUGGAU----UGGACG---ACGUGGUCA-UGGAGUUAAGGCUUCUUUCAUGUCCACGUCCACGUUCAUGAUUCAAAAGUGUCGUGCAUUAUGCAAA ...((((.(((.(.((.(((((.((.----((((((---((((((.((-(((((..........))))))).)))))))...)))))....))...))))).)).))))))))... ( -32.50) >DroSec_CAF1 10222 111 - 1 GGAGGAUUGGUCGGGCUGCAUUGGAU----UGGACGUGGACGCGGUCA-UGGAGUUAAGGCUUCUUUCAUGUCCACGUCCACGUCCAUGAUUCAAAAGUGUCGUGCAUUAUGCAAA ............(.((.(((((.((.----((((((((((((.((.((-(((((..........))))))).)).))))))))))))....))...))))).)).).......... ( -42.80) >DroSim_CAF1 11220 111 - 1 GGAGGAUUGGUCGGGCUGCAUUGGAU----UGGACGUGGACGCGGUCA-UGGAGUUAAGGCUUCUUUCAUGUCCACGUCCACGUCCAUGAUUCAAAAGUGUCGUGCAUUAUGCAAA ............(.((.(((((.((.----((((((((((((.((.((-(((((..........))))))).)).))))))))))))....))...))))).)).).......... ( -42.80) >DroEre_CAF1 9447 99 - 1 GGAUUUCUGGA------GGAUUGGUU----CGGACG---UCGUGGUCA-UGGAGUUAAGGCUUCUUUCAUGUCCACGUC---GUCCAUGAUUCAAAAGUGUCGUGCAUUAUGCAAA .(((..((...------((((((...----.(((((---.(((((.((-(((((..........))))))).))))).)---)))).))))))...)).))).(((.....))).. ( -32.40) >DroYak_CAF1 12049 98 - 1 GGA------GA------GGAUUGGUUGGGUUGGACG---ACGUGGUCAGUGGAGUUAAGGCUUCUUUCAUGUCCACGUC---GUCCAUGAUUCAAAAGUGUCGUGCAUUAUGCAAA ...------..------.(((...((((((((((((---((((((.((..(((((....))))).....)).)))))))---)))))..))))))....))).(((.....))).. ( -37.60) >consensus GGAGGAUUGGUCGGGCUGCAUUGGAU____UGGACG___ACGUGGUCA_UGGAGUUAAGGCUUCUUUCAUGUCCACGUCCACGUCCAUGAUUCAAAAGUGUCGUGCAUUAUGCAAA ....................((((((....((((((((.((((((.((..(((((....))))).....)).)))))).))))))))..))))))........(((.....))).. (-26.40 = -28.44 + 2.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:04 2006