
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,258,072 – 14,258,208 |
| Length | 136 |
| Max. P | 0.999101 |

| Location | 14,258,072 – 14,258,191 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.18 |
| Mean single sequence MFE | -40.78 |
| Consensus MFE | -28.26 |
| Energy contribution | -30.94 |
| Covariance contribution | 2.68 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.08 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979233 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14258072 119 + 22224390 UGUGUAGAGCAGGGUAGCCGGCUUUCGAUCGGCUCCAAAUUGCCACUCGUUGUAA-AGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACA (((((((.....((.(((((((....).)))))))).((((((((((........-))))))))))...(((...((((...)))).....((((....))))))).)))))))...... ( -42.10) >DroPse_CAF1 1778 90 + 1 ------------------------UCGAUCGGCUCCAAAUUGCCACUCGUUGUAA-AGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCGGCGCCUCCCC-----ACCCCCCUA ------------------------......(((....((((((((((........-))))))))))...)))......((((((((.....)))))))).......-----......... ( -29.00) >DroSim_CAF1 2473 119 + 1 UGUGUAGAGCAGGGUAGCCGGCUUUCGAUCGGCUCCAAAUUGCCACUCGUUGUAA-AGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACA (((((((.....((.(((((((....).)))))))).((((((((((........-))))))))))...(((...((((...)))).....((((....))))))).)))))))...... ( -42.10) >DroEre_CAF1 2538 119 + 1 UGUGUAGAGCAGGGGAGCCGGCUUUCGAUCGGCUCCAAAUUGCCACUCGUUGUAA-AGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACA (((((((......(((((((((....).)))))))).((((((((((........-))))))))))...(((...((((...)))).....((((....))))))).)))))))...... ( -45.90) >DroYak_CAF1 2498 120 + 1 UGUGUAUAGCAGGGGAGCUGGUUUUCGAUCGGCUCCAAAUUGCCACUCGUUGUAAAAGUGGCAAUUAACGCCACAUGACGCUGUCAACAAUUGGCCCUGGCUAGGCAUUACACACCCACA (((((((((((((((((((((((...)))))))))).....((((...(((((....(((((.......))))).((((...)))))))))))))))).)))).....))))))...... ( -44.80) >consensus UGUGUAGAGCAGGGGAGCCGGCUUUCGAUCGGCUCCAAAUUGCCACUCGUUGUAA_AGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACA (((((((.((.....((((((.........(((....((((((((((.........))))))))))...))).......(((((......))))).))))))..)).)))))))...... (-28.26 = -30.94 + 2.68)



| Location | 14,258,072 – 14,258,191 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.18 |
| Mean single sequence MFE | -43.20 |
| Consensus MFE | -29.87 |
| Energy contribution | -32.46 |
| Covariance contribution | 2.59 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.920853 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14258072 119 - 22224390 UGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACU-UUACAACGAGUGGCAAUUUGGAGCCGAUCGAAAGCCGGCUACCCUGCUCUACACA .((((((((...))))).(((.(((((((.........((((((....))))))((((((((((-(......))))))))))))))(((((..(....).))))).)))))))...))). ( -45.40) >DroPse_CAF1 1778 90 - 1 UAGGGGGGU-----GGGGAGGCGCCGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACU-UUACAACGAGUGGCAAUUUGGAGCCGAUCGA------------------------ .........-----.((..(((.(((((((....((((...)))).)))))...((((((((((-(......))))))))))).)).)))..))..------------------------ ( -34.40) >DroSim_CAF1 2473 119 - 1 UGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACU-UUACAACGAGUGGCAAUUUGGAGCCGAUCGAAAGCCGGCUACCCUGCUCUACACA .((((((((...))))).(((.(((((((.........((((((....))))))((((((((((-(......))))))))))))))(((((..(....).))))).)))))))...))). ( -45.40) >DroEre_CAF1 2538 119 - 1 UGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACU-UUACAACGAGUGGCAAUUUGGAGCCGAUCGAAAGCCGGCUCCCCUGCUCUACACA .((((((((...))))).(((.((((((((((((((((...)))..((((((.....)))))).-....)))))))))).....(((((((..(....).)))))))))))))...))). ( -50.10) >DroYak_CAF1 2498 120 - 1 UGUGGGUGUGUAAUGCCUAGCCAGGGCCAAUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACUUUUACAACGAGUGGCAAUUUGGAGCCGAUCGAAAACCAGCUCCCCUGCUAUACACA .((((((((...)))))((((.(((((((.....((((...))))..))))...(((((((((((.......))))))))))).(((((.(.........).))))))))))))..))). ( -40.70) >consensus UGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACU_UUACAACGAGUGGCAAUUUGGAGCCGAUCGAAAGCCGGCUACCCUGCUCUACACA .......(((((..((....((((.(((((((((((((...)))..((((((.....))))))......))))))))))...))))(((((.........))))).....))..))))). (-29.87 = -32.46 + 2.59)

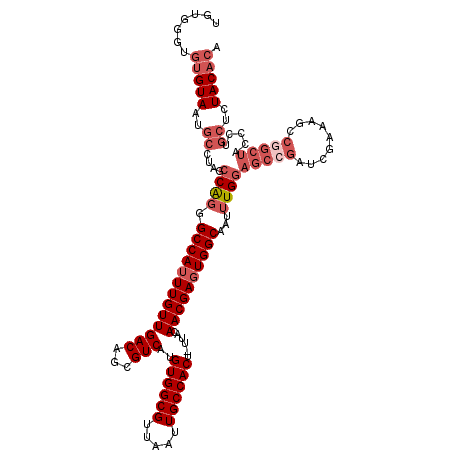
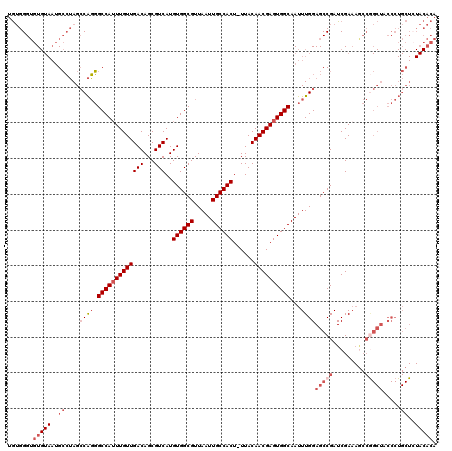
| Location | 14,258,112 – 14,258,208 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 74.22 |
| Mean single sequence MFE | -22.80 |
| Consensus MFE | -18.49 |
| Energy contribution | -17.60 |
| Covariance contribution | -0.89 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.81 |
| SVM decision value | 3.37 |
| SVM RNA-class probability | 0.999101 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14258112 96 + 22224390 UGCCACUCGUUGUAAAGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACAUUCACAUUCA-----------------CAUCUCC ((((.....((((...(((((.......))))).((((...)))))))).((((....)))))))).......................-----------------....... ( -22.30) >DroPse_CAF1 1794 108 + 1 UGCCACUCGUUGUAAAGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCGGCGCCUCCCC-----ACCCCCCUAACCCCAGCCACCUACCCGCAUUCUACAUAUGUAC (((((((........))))))).........(((((.((((((((.....)))))))).......-----...............((.........))........))))).. ( -23.80) >DroSim_CAF1 2513 96 + 1 UGCCACUCGUUGUAAAGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACACCCACAUUCA-----------------UAUCGCC ((((.....((((...(((((.......))))).((((...)))))))).((((....)))))))).......................-----------------....... ( -22.30) >consensus UGCCACUCGUUGUAAAGUGGCAAUUAACGCCACAUGACGCUGUCAACAAAUGGCCCUGGCUAGGCAUUACACACCCACAACCACAUUCA_________________UAUCUCC .((((....((((...(((((.......))))).((((...)))))))).))))((......))................................................. (-18.49 = -17.60 + -0.89)



| Location | 14,258,112 – 14,258,208 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 74.22 |
| Mean single sequence MFE | -31.33 |
| Consensus MFE | -25.95 |
| Energy contribution | -25.07 |
| Covariance contribution | -0.89 |
| Combinations/Pair | 1.10 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.775663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14258112 96 - 22224390 GGAGAUG-----------------UGAAUGUGAAUGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACUUUACAACGAGUGGCA .......-----------------............(((((((...)))))))......(((((((((((((...)))..((((((.....)))))).....)))))))))). ( -28.20) >DroPse_CAF1 1794 108 - 1 GUACAUAUGUAGAAUGCGGGUAGGUGGCUGGGGUUAGGGGGGU-----GGGGAGGCGCCGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACUUUACAACGAGUGGCA .(((....))).(((.(.(((.....))).).))).....(((-----(......))))(((((((((((((...)))..((((((.....)))))).....)))))))))). ( -33.80) >DroSim_CAF1 2513 96 - 1 GGCGAUA-----------------UGAAUGUGGGUGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACUUUACAACGAGUGGCA .......-----------------........(((.(((((((...))))))))))...(((((((((((((...)))..((((((.....)))))).....)))))))))). ( -32.00) >consensus GGAGAUA_________________UGAAUGUGGAUGUGGGUGUGUAAUGCCUAGCCAGGGCCAUUUGUUGACAGCGUCAUGUGGCGUUAAUUGCCACUUUACAACGAGUGGCA .................................................((......))(((((((((((((...)))..((((((.....)))))).....)))))))))). (-25.95 = -25.07 + -0.89)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:49:11 2006