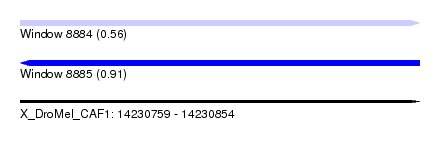

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,230,759 – 14,230,854 |
| Length | 95 |
| Max. P | 0.907456 |

| Location | 14,230,759 – 14,230,854 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 87.89 |
| Mean single sequence MFE | -22.23 |
| Consensus MFE | -19.50 |
| Energy contribution | -20.00 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558158 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14230759 95 + 22224390 UAAAUUCGCAUGUGUGCAUGCAUUCAGCUCGGGAAUUGUCACCUGAAUGUGGAAAAUACAUGCACGUACAUAUAUGU--GUAUAGAUUAGUUGCUUA .......((((((((...(((((((((.....((....))..)))))))))....))))))))..((((((....))--)))).............. ( -24.60) >DroSec_CAF1 139279 81 + 1 UAAAUUCGCAUGUGAGCAUGCAUUCAGCUCGGGAAUUGUCACCUGAAUGUGGAAAAUACAUGCACGU----------------AGAUUAGUUGCUAA .......(((((((....(((((((((.....((....))..))))))))).....)))))))..((----------------((.....))))... ( -18.40) >DroSim_CAF1 119416 97 + 1 UAAAUUCGCAUGUGUGCAUGCAUUCAGCUCGGGAAUUGUCACCUGAAUGUGGAAAAUACAUGCACGUACAUAUAUGUUCGUAUAGAUUAGUUGCUUA .......((((((((...(((((((((.....((....))..)))))))))....))))))))(((.(((....))).)))................ ( -23.80) >DroEre_CAF1 126901 89 + 1 UAAAUUCGCAUGUGCGCAUGCAUUCAGCUCGGGAAUUGUCACCUGAAUGUGGAAAAUACAUGCACGAGCAU--------AUAUAGAUUAGUUGCUUA .......(((((((....(((((((((.....((....))..))))))))).....)))))))..(((((.--------.((.....))..))))). ( -22.10) >consensus UAAAUUCGCAUGUGUGCAUGCAUUCAGCUCGGGAAUUGUCACCUGAAUGUGGAAAAUACAUGCACGUACAU________GUAUAGAUUAGUUGCUUA .......((((((((...(((((((((.....((....))..)))))))))....)))))))).................................. (-19.50 = -20.00 + 0.50)

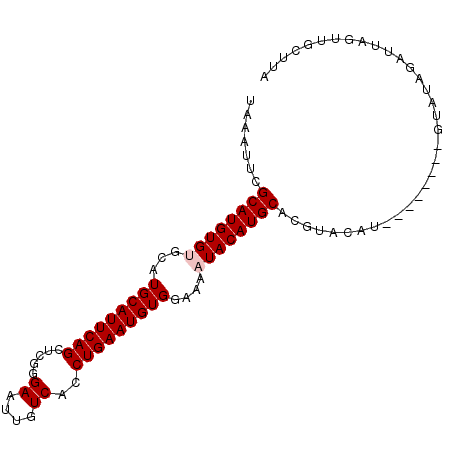
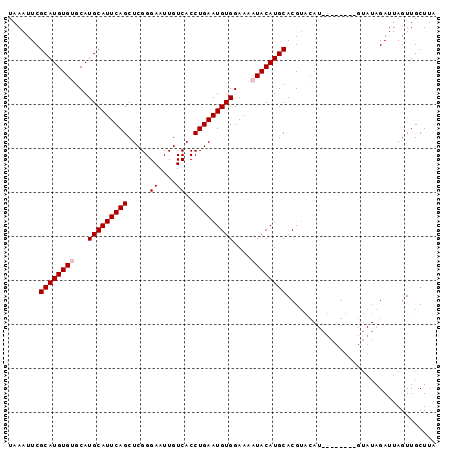
| Location | 14,230,759 – 14,230,854 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 87.89 |
| Mean single sequence MFE | -20.20 |
| Consensus MFE | -17.30 |
| Energy contribution | -17.43 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.907456 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14230759 95 - 22224390 UAAGCAACUAAUCUAUAC--ACAUAUAUGUACGUGCAUGUAUUUUCCACAUUCAGGUGACAAUUCCCGAGCUGAAUGCAUGCACACAUGCGAAUUUA ...(((.........(((--(......)))).((((((((((((......(((.((........)).)))..))))))))))))...)))....... ( -22.60) >DroSec_CAF1 139279 81 - 1 UUAGCAACUAAUCU----------------ACGUGCAUGUAUUUUCCACAUUCAGGUGACAAUUCCCGAGCUGAAUGCAUGCUCACAUGCGAAUUUA ..............----------------...(((((((......(((......))).........((((((....)).)))))))))))...... ( -16.40) >DroSim_CAF1 119416 97 - 1 UAAGCAACUAAUCUAUACGAACAUAUAUGUACGUGCAUGUAUUUUCCACAUUCAGGUGACAAUUCCCGAGCUGAAUGCAUGCACACAUGCGAAUUUA ...(((............(.(((....))).)((((((((((((......(((.((........)).)))..))))))))))))...)))....... ( -21.90) >DroEre_CAF1 126901 89 - 1 UAAGCAACUAAUCUAUAU--------AUGCUCGUGCAUGUAUUUUCCACAUUCAGGUGACAAUUCCCGAGCUGAAUGCAUGCGCACAUGCGAAUUUA ...(((........((((--------((((....))))))))......(((((((.((........))..)))))))..)))((....))....... ( -19.90) >consensus UAAGCAACUAAUCUAUAC________AUGUACGUGCAUGUAUUUUCCACAUUCAGGUGACAAUUCCCGAGCUGAAUGCAUGCACACAUGCGAAUUUA ................................((((((((((((......(((.((........)).)))..))))))))))))............. (-17.30 = -17.43 + 0.13)

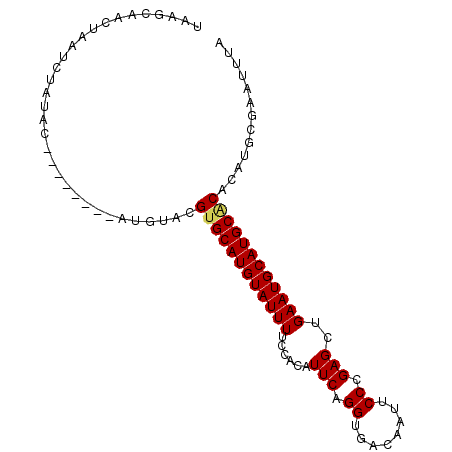
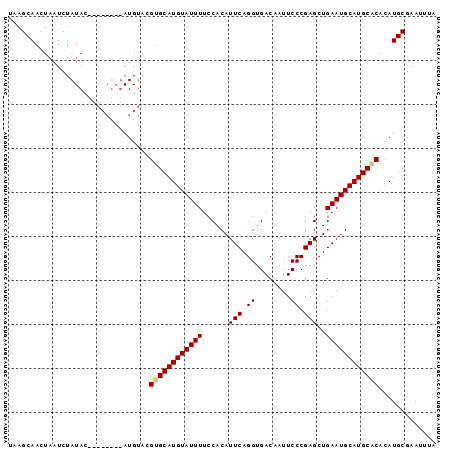
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:48:52 2006