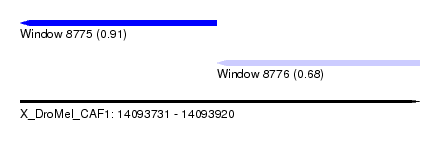

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,093,731 – 14,093,920 |
| Length | 189 |
| Max. P | 0.910205 |

| Location | 14,093,731 – 14,093,824 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 88.33 |
| Mean single sequence MFE | -22.64 |
| Consensus MFE | -18.74 |
| Energy contribution | -19.62 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.910205 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
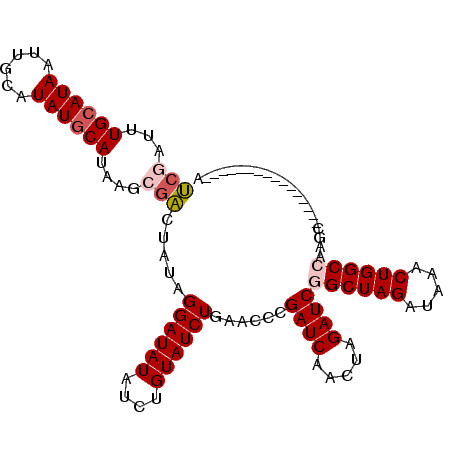
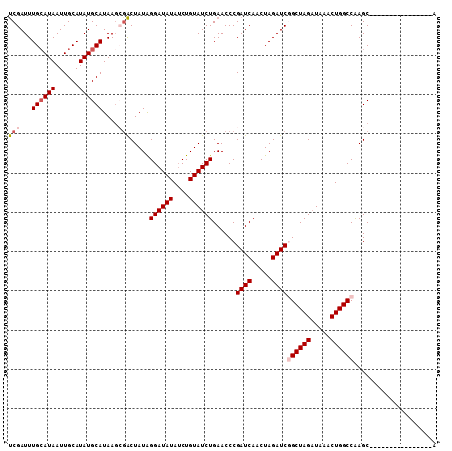
>X_DroMel_CAF1 14093731 93 - 22224390 UCGAUUUGAAUAAUUGCAUAUGCAUAAGCGACUAUAGGAUAUAUCUGUAUCUGAACCCGAUCAACUAGAUCGGCUAGAUAAACUGGCCAAGC----------------A ..............(((....)))...((.......((((((....))))))......((((.....))))((((((.....))))))..))----------------. ( -21.10) >DroSec_CAF1 4299 93 - 1 UCGAUUUGCAUAAUUGCAUAUGCAUAAGCGACUAUAGGAUAUAUCUGUAUCUGAACCCGAUCAACUAGAUCGGCUAGAUAAACUGGCCAAGC----------------A (((...((((((......))))))....))).....((((((....))))))......((((.....))))((((((.....))))))....----------------. ( -25.00) >DroSim_CAF1 3919 93 - 1 UCGAUUUGCAUAAUUGCAUAUGCAUAAGCGACUAUAGGAUAUAUCUGUAUCUGAACCCGAUCAACUAGAUCGGCUAGAUAAACUGGCCGAGC----------------A (((...((((((......))))))....))).....((((((....))))))......((((.....))))((((((.....))))))....----------------. ( -25.00) >DroEre_CAF1 4138 92 - 1 CCGAUUUGCAUAAUUGCAUAUGCAUAA-AGGCUAUAGGAUAUAUCUGUAUCUGAACCCGAUCAACUAGAUCAGCUAGCUAAACUGGCCAAGC----------------U ......((((((......))))))...-.((((...((((((....))))))......((((.....)))).(((((.....)))))..)))----------------) ( -20.70) >DroYak_CAF1 4660 109 - 1 UCGAUUUGCAUAAUUGCAUAUGCAUAAGAUACUAUGGGAUAUAGCUGUAUCUGAACCCGAUCAACUAGAUCAGCUAGAUAAACUGGCCAAGCAGACCAACACACCACCA ((....((((((......))))))...)).....(((....((((((.(((((............)))))))))))......(((......))).)))........... ( -21.40) >consensus UCGAUUUGCAUAAUUGCAUAUGCAUAAGCGACUAUAGGAUAUAUCUGUAUCUGAACCCGAUCAACUAGAUCGGCUAGAUAAACUGGCCAAGC________________A (((...((((((......))))))....))).....((((((....))))))......((((.....))))((((((.....))))))..................... (-18.74 = -19.62 + 0.88)

| Location | 14,093,824 – 14,093,920 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 93.12 |
| Mean single sequence MFE | -22.13 |
| Consensus MFE | -18.05 |
| Energy contribution | -18.17 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.678771 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
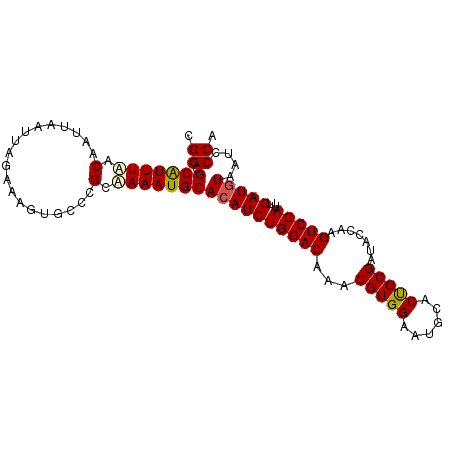
>X_DroMel_CAF1 14093824 96 - 22224390 CGGCGCAUUUAACAAUUAAUUCGAAAGUGUCCGCUAAAUGCACAUCUGGACAAACGUGGAAUGCACUGCGAUACCAAGUCCAUAGAUGUAAUCCCA .((.(((((((..........(....).......)))))))((((((((((...((..(......)..)).......))))..))))))....)). ( -21.13) >DroSec_CAF1 4392 96 - 1 CGGCGCAUUUAACAAUUAAUUAGAAAGUGCCCGCAAAAUGCACAUCUGGACAAACGUGGAAUGCACUGCGAUACCAAGUCCAUUGAUGUAAUCCCA .((.((((((..............))))))))((.....))((((((((((...((..(......)..)).......)))))..)))))....... ( -21.44) >DroSim_CAF1 4012 96 - 1 CGGCGCAUUUAACAAUUAAUUAGAAAGUGCCCGCAAAAUGCACAUCUGGACAAACGUGGAAUGCACUGCGAUGCCAAGUCCAUUGAUGUAAUCCCA .((.((((((..............))))))))((.....))((((((((((.....(((..(((...)))...))).)))))..)))))....... ( -22.44) >DroEre_CAF1 4230 96 - 1 CGGCGCGUUUUACAAUUAAUUAGAAAGUAUCCGUAAAAUGCACAUCUGGACAAACGUGGAAUGCACCGCGAUACCAAGUCCAUUGAUGUAAUCCCA .((.(((((((((...................)))))))))((((((((((...(((((......))))).......)))))..)))))....)). ( -25.91) >DroYak_CAF1 4769 94 - 1 CGGCGCAUUUUACAUUUAAUUAGAAAGUA--CGUAAAAUGCACAUCUGGACAAACGUGGAAUGCACUGCGACACCAAGUCCAUUGAUAUAAUCCCA .((.(((((((((................--.)))))))))..((((((((....(((...(((...))).)))...)))))..)))......)). ( -19.73) >consensus CGGCGCAUUUAACAAUUAAUUAGAAAGUGCCCGCAAAAUGCACAUCUGGACAAACGUGGAAUGCACUGCGAUACCAAGUCCAUUGAUGUAAUCCCA .((.(((((((.(...................).)))))))((((((((((...(((((......))))).......)))))..)))))....)). (-18.05 = -18.17 + 0.12)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:47:15 2006