
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,055,270 – 14,055,360 |
| Length | 90 |
| Max. P | 0.646731 |

| Location | 14,055,270 – 14,055,360 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 74.25 |
| Mean single sequence MFE | -18.02 |
| Consensus MFE | -3.00 |
| Energy contribution | -3.24 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.17 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.563726 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
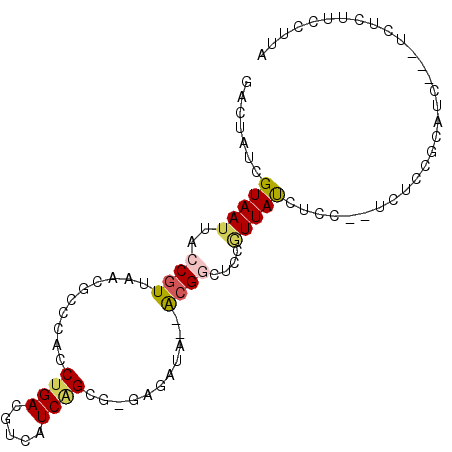
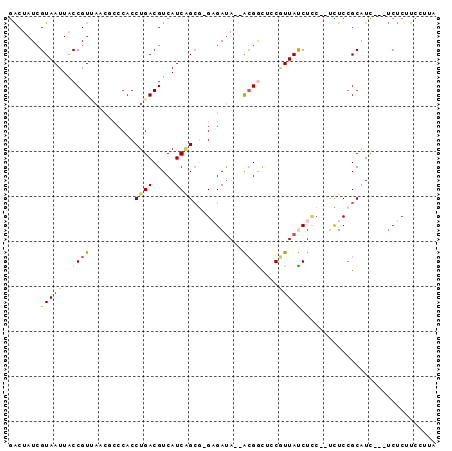
>X_DroMel_CAF1 14055270 90 + 22224390 GACUAUCGUAAUUACCGUUAACGCCCACCUGACGUCAUCGGAA-GCGAUA--GCGGCACCGUUACCUUC--UCUCCGCAUCUUCUCUCUUCCGUA .......(((((..((((((.(((....((((.....))))..-))).))--))))....)))))....--........................ ( -15.40) >DroSec_CAF1 23924 86 + 1 GACUAUCGUAAUUACCGU-AAUUACCACCUGACGUCAUCGGCG-GAGAUA--GCGGCACCGUUACCUUC--ACUCCGCAUC---UCUCCUCCCUA (((....(((((((...)-))))))(....)..)))...((.(-((((..--((((.............--...))))...---))))).))... ( -17.59) >DroEre_CAF1 24248 87 + 1 GACUAUGGUAAUUACCGUUAACUCCCACCUGACGUCAUCAGCG-GAGACA--ACGGCUGCAUUAUCUCC--UCUCCGCAUC---UCUCUUCCUUA ......(((((((((((((..((((...((((.....)))).)-)))..)--)))).)).))))))...--..........---........... ( -17.20) >DroYak_CAF1 23690 87 + 1 GACUAUCGUAAUUACCGUUAACGCCCACCUGACGUCAUCAGCG-GAGAUA--ACGACUCCGUUAUCUCC--UUUCCGCAUC---UCUCUUCCUUA ......................((....((((.....)))).(-((((((--(((....))))))))))--.....))...---........... ( -19.20) >DroAna_CAF1 19638 80 + 1 AGCUAUCGUAAUUGCCGU-------CACCUGACGUCAUCUGCAGGUGAUAUCAAGAUUGCUUUAUCUGCGGCUUCAGCGUU---ACACUU----- ......(((....(((((-------((((((.((.....))))))))))....((((......))))..)))....)))..---......----- ( -20.70) >consensus GACUAUCGUAAUUACCGUUAACGCCCACCUGACGUCAUCAGCG_GAGAUA__ACGGCUCCGUUAUCUCC__UCUCCGCAUC___UCUCUUCCUUA .......(((((..((((..........((((.....))))...........))))....))))).............................. ( -3.00 = -3.24 + 0.24)

| Location | 14,055,270 – 14,055,360 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 74.25 |
| Mean single sequence MFE | -24.72 |
| Consensus MFE | -5.06 |
| Energy contribution | -4.18 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.20 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.646731 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
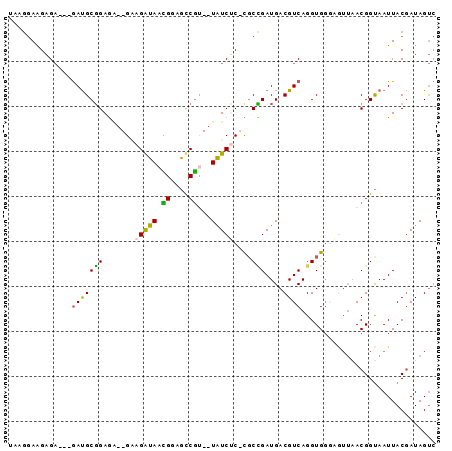
>X_DroMel_CAF1 14055270 90 - 22224390 UACGGAAGAGAGAAGAUGCGGAGA--GAAGGUAACGGUGCCGC--UAUCGC-UUCCGAUGACGUCAGGUGGGCGUUAACGGUAAUUACGAUAGUC ..((((((.((......((((...--(..(....)..).))))--..)).)-))))).(((((((.....))))))).((.......))...... ( -24.70) >DroSec_CAF1 23924 86 - 1 UAGGGAGGAGA---GAUGCGGAGU--GAAGGUAACGGUGCCGC--UAUCUC-CGCCGAUGACGUCAGGUGGUAAUU-ACGGUAAUUACGAUAGUC ...((.(((((---...((((.((--.......))....))))--..))))-).))...(((.....((.((((((-(...))))))).)).))) ( -25.80) >DroEre_CAF1 24248 87 - 1 UAAGGAAGAGA---GAUGCGGAGA--GGAGAUAAUGCAGCCGU--UGUCUC-CGCUGAUGACGUCAGGUGGGAGUUAACGGUAAUUACCAUAGUC ...........---((((((.((.--((((((((((....)))--))))))-).))..)).))))..((((.(((((....))))).)))).... ( -24.60) >DroYak_CAF1 23690 87 - 1 UAAGGAAGAGA---GAUGCGGAAA--GGAGAUAACGGAGUCGU--UAUCUC-CGCUGAUGACGUCAGGUGGGCGUUAACGGUAAUUACGAUAGUC ...........---(((.((....--((((((((((....)))--))))))-)((((.(((((((.....))))))).)))).....))...))) ( -30.10) >DroAna_CAF1 19638 80 - 1 -----AAGUGU---AACGCUGAAGCCGCAGAUAAAGCAAUCUUGAUAUCACCUGCAGAUGACGUCAGGUG-------ACGGCAAUUACGAUAGCU -----.((((.---..))))...((((.((((......)))).....((((((((.......).))))))-------)))))............. ( -18.40) >consensus UAAGGAAGAGA___GAUGCGGAGA__GAAGAUAACGGAGCCGU__UAUCUC_CGCCGAUGACGUCAGGUGGGAGUUAACGGUAAUUACGAUAGUC ..............(((((((.......(((((.((....))...)))))....)))....)))).............................. ( -5.06 = -4.18 + -0.88)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:47:07 2006