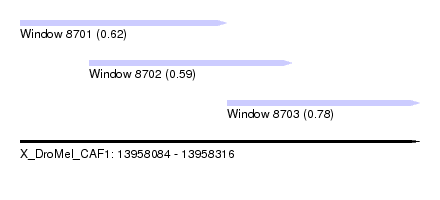
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,958,084 – 13,958,316 |
| Length | 232 |
| Max. P | 0.776559 |

| Location | 13,958,084 – 13,958,204 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.76 |
| Mean single sequence MFE | -28.13 |
| Consensus MFE | -24.70 |
| Energy contribution | -24.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.615036 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13958084 120 + 22224390 GGUCUUUGUGGCGACCAUGAAAAACGUGACAUUUUUAUGUUGCAAACUUUAAAUCUUUCAUAUCACAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGGCUGCAACAACAAC ((((........)))).........(..((((....))))..)............((((((........))))))(((((...((((.(((.((((....))))))).)))).))))).. ( -29.80) >DroSec_CAF1 9255 117 + 1 GGUCUUUGUGGCGACCAUGAAAAACGUGACAUUUUUAUGUUGCAAACUUUAAAUCUUUCAUAUCAGAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGG---CAACAACAAC ((((........)))).........(..((((....))))..)............((((((........))))))((((((..(((..(((....)))..)))...)---)))))..... ( -27.30) >DroSim_CAF1 9862 117 + 1 GGUCUUUGUGGCGACCAUGAAAAACGUGACAUUUUUAUGUUGCAAACUUUAAAUCUUUCAUAUCAGAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGG---CAACAACAAC ((((........)))).........(..((((....))))..)............((((((........))))))((((((..(((..(((....)))..)))...)---)))))..... ( -27.30) >consensus GGUCUUUGUGGCGACCAUGAAAAACGUGACAUUUUUAUGUUGCAAACUUUAAAUCUUUCAUAUCAGAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGG___CAACAACAAC ((((........)))).........(..((((....))))..)............((((((........))))))((((((..((((......))))...)))))).............. (-24.70 = -24.70 + -0.00)



| Location | 13,958,124 – 13,958,242 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 94.24 |
| Mean single sequence MFE | -23.37 |
| Consensus MFE | -17.93 |
| Energy contribution | -18.27 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.586948 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13958124 118 + 22224390 UGCAAACUUUAAAUCUUUCAUAUCACAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGGCUGCAACAACAACGUUGGCAUUUUCAGCCAACAACAACAACAACCAUUCAU ...............((((((........))))))(((((...((((.(((.((((....))))))).))))........((((((.......))))))..)))))............ ( -27.70) >DroSec_CAF1 9295 111 + 1 UGCAAACUUUAAAUCUUUCAUAUCAGAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGG---CAACAACAACGUUGGCAUUUUCAGU----GACAACAACAACCAUUCAU ...............((((......))))(((((.((((((..(((..(((....)))..)))...)---))))).....((((.(((.....))----).))))........))))) ( -21.20) >DroSim_CAF1 9902 111 + 1 UGCAAACUUUAAAUCUUUCAUAUCAGAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGG---CAACAACAACGUUGGCAUUUUCAGU----GACAACAACAACCAUUCAU ...............((((......))))(((((.((((((..(((..(((....)))..)))...)---))))).....((((.(((.....))----).))))........))))) ( -21.20) >consensus UGCAAACUUUAAAUCUUUCAUAUCAGAAAAUGAAAUGUUGCAAUGCAAUUGACUGCAACAGCAGCGG___CAACAACAACGUUGGCAUUUUCAGU____GACAACAACAACCAUUCAU .........................(((((((...((((((..((((......))))...))))))....((((......)))).))))))).......................... (-17.93 = -18.27 + 0.33)



| Location | 13,958,204 – 13,958,316 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 93.07 |
| Mean single sequence MFE | -31.97 |
| Consensus MFE | -26.08 |
| Energy contribution | -26.53 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.776559 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13958204 112 + 22224390 GUUGGCAUUUUCAGCCAACAACAACAACAACCAUUCAUGUGCACAUGGUUCUCAGUUCUUUGGCAUCUUUUUGUUGUCUUUGAUUGGGAUUUUUGUGUGCUGGUGGCUUUGC ((((((.......))))))...........((((.((.(..((((....((((((((....((((.(.....).))))...))))))))....))))..)))))))...... ( -34.20) >DroSec_CAF1 9372 107 + 1 GUUGGCAUUUUCAGU----GACAACAACAACCAUUCAUGUGCACAUG-UGCCCAGUUCUUUGGCAUCUUUUUGUUGUCUUUGAUUAGGAUUUUUGUGUGCUGGUGGCUUUGC ((((.(((.....))----).)))).....((((.((.(..((((..-..((.((((....((((.(.....).))))...)))).)).....))))..)))))))...... ( -24.60) >DroSim_CAF1 9979 108 + 1 GUUGGCAUUUUCAGU----GACAACAACAACCAUUCAUGUGCACAUGCUUCCCAGUUCUUUGGCAGCUUUUUGUUGUCUUUGAUUGGGAUUUUUGUGUGCUGGUGGCUUUGC ((((.(((.....))----).)))).....((((.((.(..((((....((((((((....((((((.....))))))...))))))))....))))..)))))))...... ( -37.10) >consensus GUUGGCAUUUUCAGU____GACAACAACAACCAUUCAUGUGCACAUG_UUCCCAGUUCUUUGGCAUCUUUUUGUUGUCUUUGAUUGGGAUUUUUGUGUGCUGGUGGCUUUGC ((((.................)))).....((((.((.(..((((....((((((((....((((.(.....).))))...))))))))....))))..)))))))...... (-26.08 = -26.53 + 0.45)


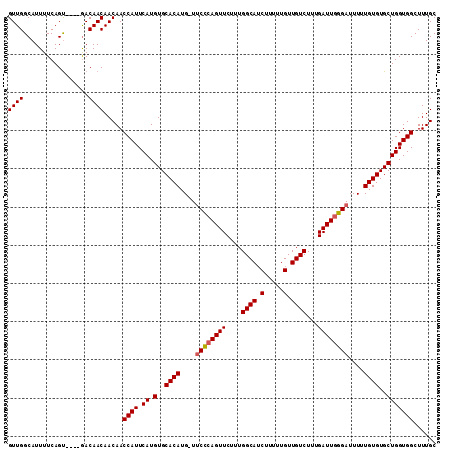
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:09 2006