
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,954,658 – 13,954,767 |
| Length | 109 |
| Max. P | 0.849850 |

| Location | 13,954,658 – 13,954,767 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 86.59 |
| Mean single sequence MFE | -41.10 |
| Consensus MFE | -30.99 |
| Energy contribution | -31.17 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849850 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13954658 109 + 22224390 G--GGGGGGGUGGUGCUAGGUGAGGAGCCGGCGAAUGGAAUUGCCAACU-------GACAACGUGGGCGGCAACAUCUCGCACAGUUGGCAACAUUCUAUUUUCCAAUUCUUCCGAAG (--((((((.(((.(((.(((.....))))))(((((((((((((((((-------(.....(((((.(....)..))))).)))))))))..))))))))).))).))))))).... ( -42.40) >DroSec_CAF1 6101 110 + 1 GU-UGGGGGGAGGGGUGAGGCGUGGAGCCGGCGAAUGGAAUUGCCAACU-------GACAACGUGGGCGGCAACAUCUCGCACAGUUGGCAACAUUCUAUUUUCCAAUUCUUUCGAAG .(-((..((((.......(((.....)))((.(((((((((((((((((-------(.....(((((.(....)..))))).)))))))))..))))))))).))..))))..))).. ( -42.80) >DroSim_CAF1 6503 111 + 1 GUUUGGGGGGAGGGGUGAGGCGUGGAGCCGGCGAAUGGAAUUGCCAACU-------GACAACGUGGGCGGCAACAUCUCGCACAGUUGGCAACAUUCUAUUUUCCAAUUCUUUCGAAG .((((..((((.......(((.....)))((.(((((((((((((((((-------(.....(((((.(....)..))))).)))))))))..))))))))).))..))))..)))). ( -43.70) >DroYak_CAF1 6539 118 + 1 GGAUGGCAAAGGGGGGGAUACGGGGAGCUGGCAAAUGAAAUUGCCAACUGACAACUGACAACGUGGGCGGCAACAUCUUGCACAGUUGGCAACAUUCUAUUUUCCAAUUCUUCCGAAG ...........((((((((...(((((.(((..((((...((((((((((...........((....))((((....)))).))))))))))))))))).))))).)))))))).... ( -35.50) >consensus GU_UGGGGGGAGGGGUGAGGCGUGGAGCCGGCGAAUGGAAUUGCCAACU_______GACAACGUGGGCGGCAACAUCUCGCACAGUUGGCAACAUUCUAUUUUCCAAUUCUUCCGAAG ....(((((((.......(((.....)))((.(((((((((((((((((.......(....)(((((.(....)..)))))..))))))))..))))))))).))..))))))).... (-30.99 = -31.17 + 0.19)

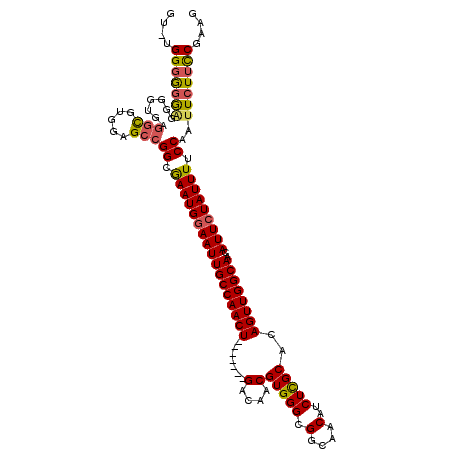

| Location | 13,954,658 – 13,954,767 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 86.59 |
| Mean single sequence MFE | -31.77 |
| Consensus MFE | -23.23 |
| Energy contribution | -22.85 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.570180 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13954658 109 - 22224390 CUUCGGAAGAAUUGGAAAAUAGAAUGUUGCCAACUGUGCGAGAUGUUGCCGCCCACGUUGUC-------AGUUGGCAAUUCCAUUCGCCGGCUCCUCACCUAGCACCACCCCCCC--C .(((....))).(((......(((((((((((((((.((((..((........))..)))))-------)))))))))...)))))((..(........)..)).))).......--. ( -30.10) >DroSec_CAF1 6101 110 - 1 CUUCGAAAGAAUUGGAAAAUAGAAUGUUGCCAACUGUGCGAGAUGUUGCCGCCCACGUUGUC-------AGUUGGCAAUUCCAUUCGCCGGCUCCACGCCUCACCCCUCCCCCCA-AC .(((....)))((((......(((((((((((((((.((((..((........))..)))))-------)))))))))...)))))...(((.....)))............)))-). ( -32.90) >DroSim_CAF1 6503 111 - 1 CUUCGAAAGAAUUGGAAAAUAGAAUGUUGCCAACUGUGCGAGAUGUUGCCGCCCACGUUGUC-------AGUUGGCAAUUCCAUUCGCCGGCUCCACGCCUCACCCCUCCCCCCAAAC .(((....)))((((......(((((((((((((((.((((..((........))..)))))-------)))))))))...)))))...(((.....)))............)))).. ( -33.20) >DroYak_CAF1 6539 118 - 1 CUUCGGAAGAAUUGGAAAAUAGAAUGUUGCCAACUGUGCAAGAUGUUGCCGCCCACGUUGUCAGUUGUCAGUUGGCAAUUUCAUUUGCCAGCUCCCCGUAUCCCCCCCUUUGCCAUCC ...(((..((.((((.((((.(((.(((((((((((.((((..((.(..((....))..).)).)))))))))))))))))))))).)))).)).))).................... ( -30.90) >consensus CUUCGAAAGAAUUGGAAAAUAGAAUGUUGCCAACUGUGCGAGAUGUUGCCGCCCACGUUGUC_______AGUUGGCAAUUCCAUUCGCCGGCUCCACGCCUCACCCCUCCCCCCA_AC .(((....)))..(((.....((((((((((((((..((((....))))((....))............)))))))))...)))))......)))....................... (-23.23 = -22.85 + -0.38)

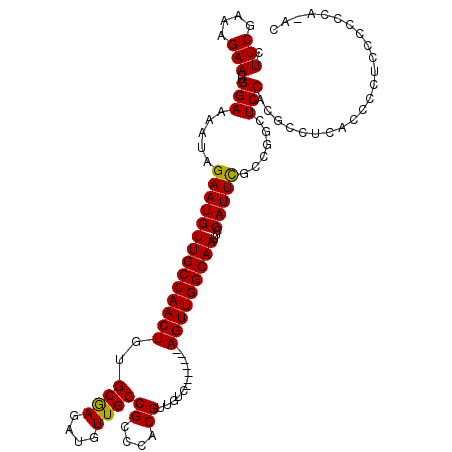

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:05 2006