
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,951,963 – 13,952,081 |
| Length | 118 |
| Max. P | 0.858771 |

| Location | 13,951,963 – 13,952,081 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.84 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -25.19 |
| Energy contribution | -24.97 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.858771 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13951963 118 + 22224390 --CUUUAGCACACACAGCUGUUGAAAUAAACUUUCUCGCUUUGGGAAAUCUAUUUCUAAUCUAUUUGUUUGCCACUCAAGUGGGAACGCAAGGCCCCCCCUCUGUCCAUCCCUUCAGCUG --............((((((..((((((...((((((.....))))))..))))))..........((((.((((....))))))))((((((.....))).))).........)))))) ( -25.00) >DroSec_CAF1 3419 117 + 1 UUUUUUAGC--ACACAGCUGUUGAAAUAAACUUUCUCACUUUGGGAAAUCUAUUUCUAAUCUAUUUGUUUGCCACUCAAGUGGGAUGGCAAGGCCC-CCCUCUGUCCAUCCCUUCAGCUG .........--...((((((..((((((...((((((.....))))))..))))))......(((((.........)))))((((((((((((...-.))).)).)))))))..)))))) ( -32.60) >DroSim_CAF1 3814 117 + 1 UUUUUUAGC--GCACAGCUGUUGAAAUAAACUUUCUCACUUUGGGAAAUCUAUUUCUAAUCUAUUUGUUUGCCACUCAAGUGGGAUCACAAGGCCC-CCCUCUGUCCAUCCCUUCAGCUG .........--...((((((..((((((...((((((.....))))))..))))))......(((((.........)))))(((((.((((((...-.))).)))..)))))..)))))) ( -25.80) >consensus UUUUUUAGC__ACACAGCUGUUGAAAUAAACUUUCUCACUUUGGGAAAUCUAUUUCUAAUCUAUUUGUUUGCCACUCAAGUGGGAUCGCAAGGCCC_CCCUCUGUCCAUCCCUUCAGCUG ..............((((((..((((((...((((((.....))))))..))))))......(((((.........)))))((((..((((((.....))).)))...))))..)))))) (-25.19 = -24.97 + -0.22)

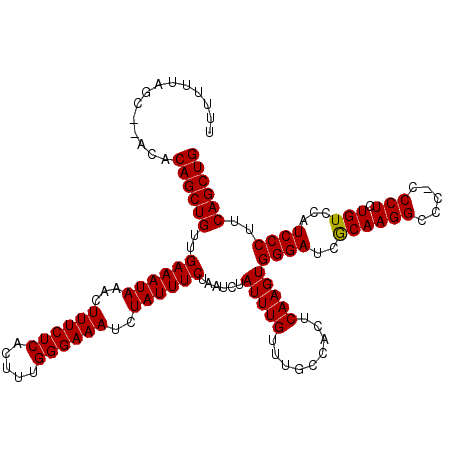

| Location | 13,951,963 – 13,952,081 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.84 |
| Mean single sequence MFE | -33.07 |
| Consensus MFE | -29.19 |
| Energy contribution | -29.30 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.763706 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13951963 118 - 22224390 CAGCUGAAGGGAUGGACAGAGGGGGGGCCUUGCGUUCCCACUUGAGUGGCAAACAAAUAGAUUAGAAAUAGAUUUCCCAAAGCGAGAAAGUUUAUUUCAACAGCUGUGUGUGCUAAAG-- ((((((..(..((((((...(((((.((...)).))))).((((..(((.((((.....).(((....))).))).)))...))))...))))))..)..))))))............-- ( -32.90) >DroSec_CAF1 3419 117 - 1 CAGCUGAAGGGAUGGACAGAGGG-GGGCCUUGCCAUCCCACUUGAGUGGCAAACAAAUAGAUUAGAAAUAGAUUUCCCAAAGUGAGAAAGUUUAUUUCAACAGCUGUGU--GCUAAAAAA ((((((..(((((((...((((.-...)))).))))))).........................(((((((((((.((.....).).)))))))))))..))))))...--......... ( -36.00) >DroSim_CAF1 3814 117 - 1 CAGCUGAAGGGAUGGACAGAGGG-GGGCCUUGUGAUCCCACUUGAGUGGCAAACAAAUAGAUUAGAAAUAGAUUUCCCAAAGUGAGAAAGUUUAUUUCAACAGCUGUGC--GCUAAAAAA ((((((..(((((.(...((((.-...)))).).))))).........................(((((((((((.((.....).).)))))))))))..))))))...--......... ( -30.30) >consensus CAGCUGAAGGGAUGGACAGAGGG_GGGCCUUGCGAUCCCACUUGAGUGGCAAACAAAUAGAUUAGAAAUAGAUUUCCCAAAGUGAGAAAGUUUAUUUCAACAGCUGUGU__GCUAAAAAA ((((((..(((((.(...((((.....)))).).))))).........................(((((((((((.((.....).).)))))))))))..)))))).............. (-29.19 = -29.30 + 0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:01 2006