
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,934,553 – 13,934,654 |
| Length | 101 |
| Max. P | 0.975451 |

| Location | 13,934,553 – 13,934,654 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 82.20 |
| Mean single sequence MFE | -43.62 |
| Consensus MFE | -39.12 |
| Energy contribution | -39.50 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.905101 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
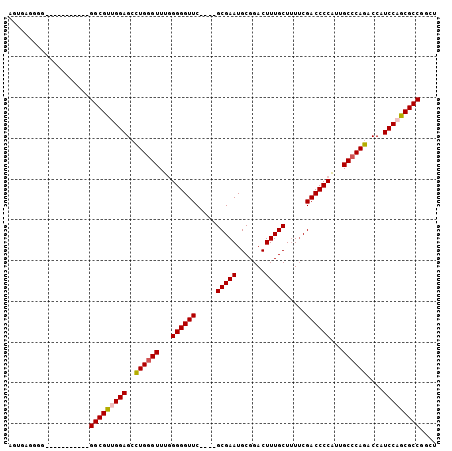
>X_DroMel_CAF1 13934553 101 + 22224390 AGUGAGGGGGGGUGGGGGAGGGCGUUGGAGCCUGGGUUUGGGGGUUC----GCGAAUGCGGACUUUGCUUUUCGACCCCAUUGCCCAGACCAUCCUGCGCCGGUU ..........((((.((((((((......))((((((.(((((((((----((....)))))).(((.....))))))))..)))))).)).)))).)))).... ( -44.00) >DroSec_CAF1 14403 94 + 1 AGUGAGGGG-----------GGCGUUGGAGCCUGGGUUUGGGGGUUCGAACGCGAAUGCGGACUUUGCUUUUCGACCCCAUUGCCCAGACCAUCCAGCGCCGGCU .........-----------(((((((((..((((((...(((((.((((.(((((.......)))))..)))))))))...))))))....))))))))).... ( -46.30) >DroSim_CAF1 20178 90 + 1 AGUGAGGGG-----------GGCGUUGGAGCCUGGGUUUGGGGGUUC----GCGAAUGCGGACUUUGCUUUUCGACCCCAUUGCCCAGACCAUCCAGCGCCGGCU .........-----------(((((((((..((((((.(((((((((----((....)))))).(((.....))))))))..))))))....))))))))).... ( -44.20) >DroEre_CAF1 15766 94 + 1 AGGGGG------CGGCAUGGGGCGCUGGAGAUUGUGUUU-GGGGUUC----GCGAAUGCGGACUUUGCUCUGCGACCCCGUUGCCCAGUCCAUCCUGCGCCGGCU ((((((------(((((((((((((.((((........(-..(((((----((....)))))))..)))))))).))))).))))..))))..)))((....)). ( -40.00) >consensus AGUGAGGGG___________GGCGUUGGAGCCUGGGUUUGGGGGUUC____GCGAAUGCGGACUUUGCUUUUCGACCCCAUUGCCCAGACCAUCCAGCGCCGGCU ....................(((((((((..((((((...((((((.....(((((.......))))).....))))))...))))))....))))))))).... (-39.12 = -39.50 + 0.38)


| Location | 13,934,553 – 13,934,654 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 82.20 |
| Mean single sequence MFE | -35.75 |
| Consensus MFE | -29.24 |
| Energy contribution | -30.11 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.75 |
| SVM RNA-class probability | 0.975451 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13934553 101 - 22224390 AACCGGCGCAGGAUGGUCUGGGCAAUGGGGUCGAAAAGCAAAGUCCGCAUUCGC----GAACCCCCAAACCCAGGCUCCAACGCCCUCCCCCACCCCCCCUCACU ....((((..(((..(((((((....(((((((..((((.......)).))..)----).)))))....))))))))))..)))).................... ( -35.00) >DroSec_CAF1 14403 94 - 1 AGCCGGCGCUGGAUGGUCUGGGCAAUGGGGUCGAAAAGCAAAGUCCGCAUUCGCGUUCGAACCCCCAAACCCAGGCUCCAACGCC-----------CCCCUCACU ....((((.((((..(((((((....(((((((((..........(((....))))))).)))))....))))))))))).))))-----------......... ( -38.80) >DroSim_CAF1 20178 90 - 1 AGCCGGCGCUGGAUGGUCUGGGCAAUGGGGUCGAAAAGCAAAGUCCGCAUUCGC----GAACCCCCAAACCCAGGCUCCAACGCC-----------CCCCUCACU ....((((.((((..(((((((....(((((((..((((.......)).))..)----).)))))....))))))))))).))))-----------......... ( -37.10) >DroEre_CAF1 15766 94 - 1 AGCCGGCGCAGGAUGGACUGGGCAACGGGGUCGCAGAGCAAAGUCCGCAUUCGC----GAACCCC-AAACACAAUCUCCAGCGCCCCAUGCCG------CCCCCU ....(((((.(((..((.(((....)((((((((..(((.......)).)..))----).)))))-.....)).))))).)))))........------...... ( -32.10) >consensus AGCCGGCGCAGGAUGGUCUGGGCAAUGGGGUCGAAAAGCAAAGUCCGCAUUCGC____GAACCCCCAAACCCAGGCUCCAACGCC___________CCCCUCACU ....((((.((((..(((((((....(((((((((..((.......)).)))).......)))))....))))))))))).)))).................... (-29.24 = -30.11 + 0.87)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:49 2006