
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,932,095 – 13,932,197 |
| Length | 102 |
| Max. P | 0.718025 |

| Location | 13,932,095 – 13,932,197 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 84.94 |
| Mean single sequence MFE | -32.48 |
| Consensus MFE | -25.56 |
| Energy contribution | -25.62 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.718025 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13932095 102 + 22224390 GCUUGGAUUUGG------AUUCCGAUUUCAGAGCCAGCUUGGGGAUCCACAUCCACAUUCACAUCCAUUCGGACGGACGUUCAUCCAUCGUGGCUGUGCCACUUCAAC ...(((((.(((------((((((....).(((....))).)))))))).)))))........(((....))).(((......)))...(((((...)))))...... ( -29.80) >DroSec_CAF1 12071 96 + 1 GCUUGGAUUUGG------AUUCGGAUUUCGGAGCCAGCUUGGGGAUCCACAUC------CACAUCCAUUCGGACGGACGUUCAUCCAUCGUGGCUGUGCCACUUUAAC (..(((((.(((------((..((((..(.(((....))).)..))))..)))------)).)))))..)((((....)))).......(((((...)))))...... ( -36.30) >DroSim_CAF1 15648 102 + 1 GCUUGGAUUUGG------AUUCGGAUUUCGGAGCCAGCUUGGGGAUCCACAUCCACAUCCACAUCCAUUCGGACGGACGUUCAUCCAUCGUGGCUGUGCCACUUCAAC ...(((((.(((------((..((((..(.(((....))).)..))))..))))).)))))..(((....))).(((......)))...(((((...)))))...... ( -36.50) >DroYak_CAF1 14238 89 + 1 GCUUGGAUUUGGAUUCGGAUUCGGAUUUCGGAGCCAGUUUGGGGAUCCA-------------------UCGGACGGACGUUCAUCCAUCGUGGCUGUGCCACUUCAAC ..(((((..(((((........((((..(.(((....))).)..)))).-------------------..((((....)))))))))..(((((...)))))))))). ( -27.30) >consensus GCUUGGAUUUGG______AUUCGGAUUUCGGAGCCAGCUUGGGGAUCCACAUC______CACAUCCAUUCGGACGGACGUUCAUCCAUCGUGGCUGUGCCACUUCAAC ...(((((..............((((..(.(((....))).)..))))......................((((....)))))))))..(((((...)))))...... (-25.56 = -25.62 + 0.06)


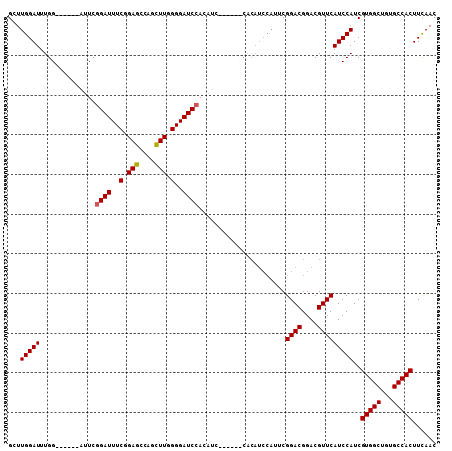
| Location | 13,932,095 – 13,932,197 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 84.94 |
| Mean single sequence MFE | -32.16 |
| Consensus MFE | -20.31 |
| Energy contribution | -20.12 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.538908 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13932095 102 - 22224390 GUUGAAGUGGCACAGCCACGAUGGAUGAACGUCCGUCCGAAUGGAUGUGAAUGUGGAUGUGGAUCCCCAAGCUGGCUCUGAAAUCGGAAU------CCAAAUCCAAGC ......(((((...))))).....((..((((((((....))))))))..)).(((((.(((((..((.....)).((((....))))))------))).)))))... ( -31.20) >DroSec_CAF1 12071 96 - 1 GUUAAAGUGGCACAGCCACGAUGGAUGAACGUCCGUCCGAAUGGAUGUG------GAUGUGGAUCCCCAAGCUGGCUCCGAAAUCCGAAU------CCAAAUCCAAGC ......(((((...)))))(((((((....)))))))....(((((.((------(((.(((((..(..((....))..)..))))).))------))).)))))... ( -35.10) >DroSim_CAF1 15648 102 - 1 GUUGAAGUGGCACAGCCACGAUGGAUGAACGUCCGUCCGAAUGGAUGUGGAUGUGGAUGUGGAUCCCCAAGCUGGCUCCGAAAUCCGAAU------CCAAAUCCAAGC ......(((((...)))))(((((((....)))))))....(((((.(((((.(((((.((((.((.......)).))))..))))).))------))).)))))... ( -40.40) >DroYak_CAF1 14238 89 - 1 GUUGAAGUGGCACAGCCACGAUGGAUGAACGUCCGUCCGA-------------------UGGAUCCCCAAACUGGCUCCGAAAUCCGAAUCCGAAUCCAAAUCCAAGC ((((..(((((...)))))(((((((....))))))))))-------------------)((((........(((...((.....))...))).......)))).... ( -21.96) >consensus GUUGAAGUGGCACAGCCACGAUGGAUGAACGUCCGUCCGAAUGGAUGUG______GAUGUGGAUCCCCAAGCUGGCUCCGAAAUCCGAAU______CCAAAUCCAAGC ......(((((...)))))(((((((....)))))))......................((((.((.......)).))))............................ (-20.31 = -20.12 + -0.19)


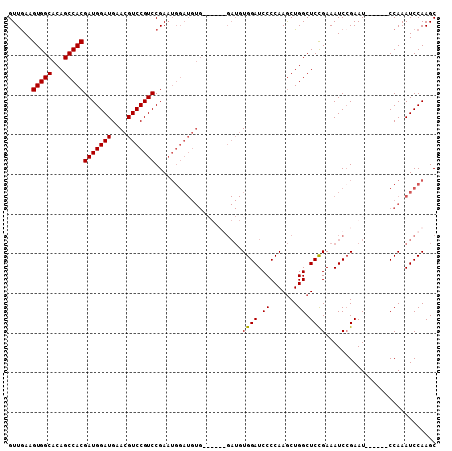
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:42 2006