
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,527,095 – 1,527,191 |
| Length | 96 |
| Max. P | 0.791337 |

| Location | 1,527,095 – 1,527,191 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 83.51 |
| Mean single sequence MFE | -26.82 |
| Consensus MFE | -16.21 |
| Energy contribution | -18.02 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.60 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521383 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
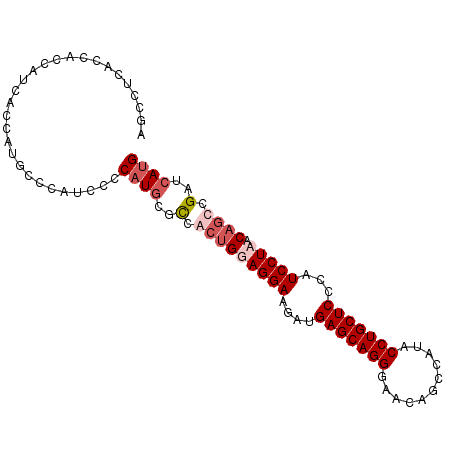
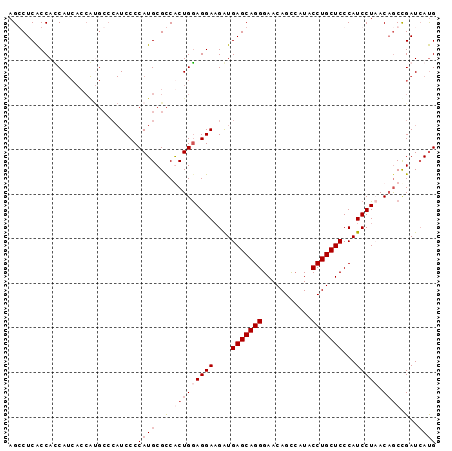
>X_DroMel_CAF1 1527095 96 + 22224390 AGGCUCGCCAUCAUCACCAUGCCCAUCCCCAUGCGCCGCUGAAGGAAGAUGAGCAGGGAACAGCAAUACCUGCUCCCAUCCUAACAACUGAUAACG .((....))(((((((.(..((.(((....))).)).).)))((((.(..(((((((...........))))))).).))))......)))).... ( -22.30) >DroSec_CAF1 125 96 + 1 AGGCUCACCAUCAUCACCAUGCCCAUCCCCAUGCGCCGCUGGAGGAAGAUGAGCAGGGAACAGCCAUACCUGCUCCCAUCCUAACAGCCGAUCAUG .(((................)))......((((..(.((((.((((.(..(((((((...........))))))).).))))..)))).)..)))) ( -27.79) >DroEre_CAF1 119 84 + 1 AGCCUCACCACCA------------UCCCCAUGCGUCACUGGAGGAUGGUGAGCAGGGAACAGUCAUACCUGCUCCCAUCCUUACUGCCGAUCAUG .............------------....((((((.((...((((((((.(((((((...........)))))))))))))))..)).))..)))) ( -30.60) >DroYak_CAF1 1 96 + 1 AGCCUCACCACCAUCACCGCGCCCAUCCCCAUGCGCCACUGGAGGAAGAUGAGCAGGGAACAGCCAUUCCUGCUCCCAUCCUUACAGCCGAUCAUG ............(((...((((..........))))..((((((((.(..(((((((((.......))))))))).).))))).)))..))).... ( -26.60) >consensus AGCCUCACCACCAUCACCAUGCCCAUCCCCAUGCGCCACUGGAGGAAGAUGAGCAGGGAACAGCCAUACCUGCUCCCAUCCUAACAGCCGAUCAUG .............................((((..(.(((((((((....(((((((...........)))))))...))))).)))).)..)))) (-16.21 = -18.02 + 1.81)

| Location | 1,527,095 – 1,527,191 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 83.51 |
| Mean single sequence MFE | -36.85 |
| Consensus MFE | -28.74 |
| Energy contribution | -28.55 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.791337 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
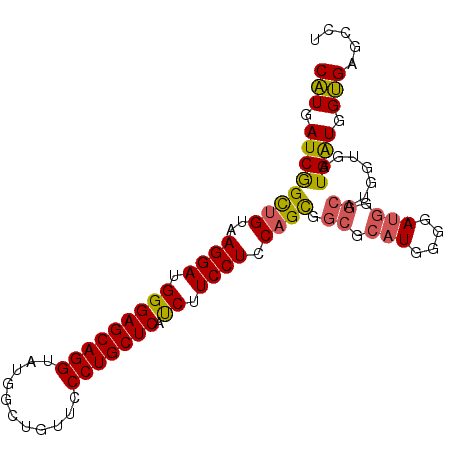
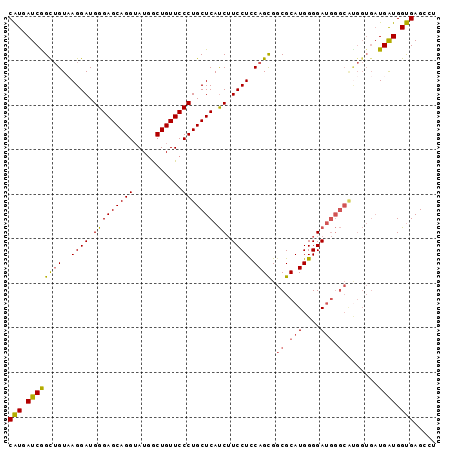
>X_DroMel_CAF1 1527095 96 - 22224390 CGUUAUCAGUUGUUAGGAUGGGAGCAGGUAUUGCUGUUCCCUGCUCAUCUUCCUUCAGCGGCGCAUGGGGAUGGGCAUGGUGAUGAUGGCGAGCCU ((((((((.(..(((....((((((((......))))))))(((((((((.((....((...))..))))))))))))))..)))))))))..... ( -37.30) >DroSec_CAF1 125 96 - 1 CAUGAUCGGCUGUUAGGAUGGGAGCAGGUAUGGCUGUUCCCUGCUCAUCUUCCUCCAGCGGCGCAUGGGGAUGGGCAUGGUGAUGAUGGUGAGCCU .......((((........((((((((......)))))))).((.((((.((..(((...((.(((....))).)).))).)).)))))).)))). ( -35.50) >DroEre_CAF1 119 84 - 1 CAUGAUCGGCAGUAAGGAUGGGAGCAGGUAUGACUGUUCCCUGCUCACCAUCCUCCAGUGACGCAUGGGGA------------UGGUGGUGAGGCU .......(((.........((((((((......))))))))..((((((((((((((........))))).------------.)))))))))))) ( -37.20) >DroYak_CAF1 1 96 - 1 CAUGAUCGGCUGUAAGGAUGGGAGCAGGAAUGGCUGUUCCCUGCUCAUCUUCCUCCAGUGGCGCAUGGGGAUGGGCGCGGUGAUGGUGGUGAGGCU (((.((((.((((......((((((((......)))))))).((((((((....(((........))))))))))))))))))).)))........ ( -37.40) >consensus CAUGAUCGGCUGUAAGGAUGGGAGCAGGUAUGGCUGUUCCCUGCUCAUCUUCCUCCAGCGGCGCAUGGGGAUGGGCAUGGUGAUGAUGGUGAGCCU (((.((((((((..((((.(((((((((...........))))))).)).)))).)))).((.(((....))).)).......)))).)))..... (-28.74 = -28.55 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:47:08 2006