
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,924,084 – 13,924,223 |
| Length | 139 |
| Max. P | 0.807354 |

| Location | 13,924,084 – 13,924,199 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.73 |
| Mean single sequence MFE | -37.63 |
| Consensus MFE | -32.80 |
| Energy contribution | -33.20 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537279 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
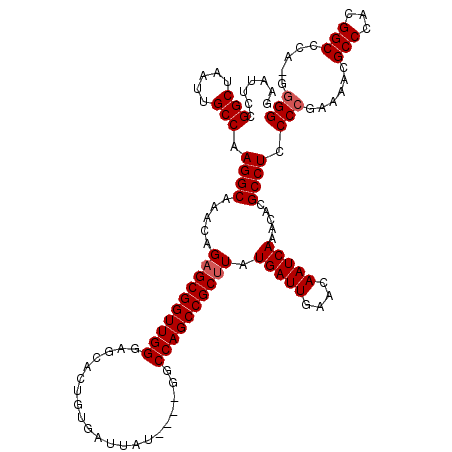
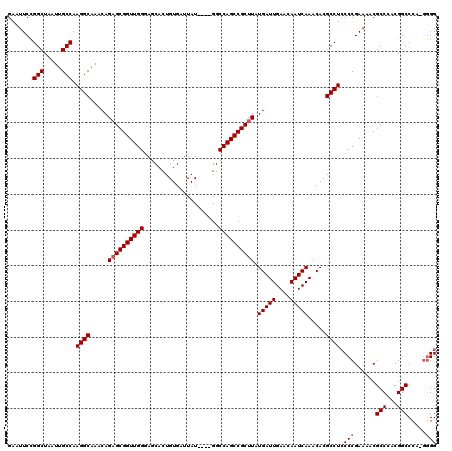
>X_DroMel_CAF1 13924084 115 - 22224390 GAAUUCUGGCUAAUUGCCAAGGCAAACAGAGCGGUUGGUAGCACUGUGAUUAU----GGCCAGCCGCUUAUGAUUGAACAAUCAAACACGCCUCCCCGAAAACGCCCACGGCCCA-GGGG ......((((.....))))((((.....(((((((((((..............----.))))))))))).(((((....))))).....))))((((......(((...)))...-)))) ( -40.26) >DroSec_CAF1 4253 114 - 1 GAAUUCCGGCUAAUUGCCAAGGCAAACAGAGCGGUUGGGAGCACUGUGAUUAU----GGCCAGCCGCUUAUGAUUGAACAAUCAAACACGCCUCCCCGAAAACGCCCACGGCCCA-G-GG .......(((.....))).((((.....((((((((((...............----..)))))))))).(((((....))))).....)))).(((......(((...)))...-)-)) ( -34.83) >DroSim_CAF1 6208 115 - 1 GAAUUCCGGCUAAUUGCCAAGGCAAACAGAGCGGUUGGGAGCACUGUGAUUAU----GGCCAGCCGCUUAUGAUUGAACAAUCAAACACGCCUCCCCGAAAACGCCCACGGCCCA-GGGG .......(((.....))).((((.....((((((((((...............----..)))))))))).(((((....))))).....))))((((......(((...)))...-)))) ( -37.93) >DroEre_CAF1 6169 115 - 1 GAAUUCUGGCUAAUUGCCAAGGCAAACAGAGCGGUUGGGAGCACUGUGAUUAU----GGCCAGCCGCUUAUGAUUGAACAAUCAAACACGCCUCCCCGAAAACGCCCACGGCCCA-GGGG ......((((.....))))((((.....((((((((((...............----..)))))))))).(((((....))))).....))))((((......(((...)))...-)))) ( -38.43) >DroYak_CAF1 5742 120 - 1 GAAUUCCGGCUAAUUGCCAAGGCAAACAGUGCGGUUGGGAGCACUGUGAUUAUGGAUGGCCAGCCGCUUAUGAUUGAACAAUCAAACACGCCUACCCGAAAACGCCCACGGCCCUCAGGG .......(((.....))).((((..(((((((........)))))))(((((((..(((....)))..)))))))..............)))).(((......(((...))).....))) ( -36.70) >consensus GAAUUCCGGCUAAUUGCCAAGGCAAACAGAGCGGUUGGGAGCACUGUGAUUAU____GGCCAGCCGCUUAUGAUUGAACAAUCAAACACGCCUCCCCGAAAACGCCCACGGCCCA_GGGG .......(((.....))).((((.....((((((((((.....................)))))))))).(((((....))))).....)))).(((......(((...))).....))) (-32.80 = -33.20 + 0.40)

| Location | 13,924,123 – 13,924,223 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 96.31 |
| Mean single sequence MFE | -26.44 |
| Consensus MFE | -25.44 |
| Energy contribution | -25.00 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.807354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
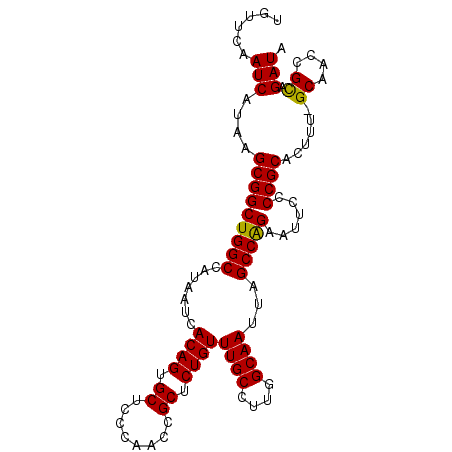
>X_DroMel_CAF1 13924123 100 + 22224390 UGUUCAAUCAUAAGCGGCUGGCCAUAAUCACAGUGCUACCAACCGCUCUGUUUGCCUUGGCAAUUAGCCAGAAUUCCCCGCACUUUGGCAAACGUAGAUA ......(((...(((((.(((.(((.......)))...))).)))))..((((((((((((.....)))))...............)))))))...))). ( -26.06) >DroSec_CAF1 4291 99 + 1 UGUUCAAUCAUAAGCGGCUGGCCAUAAUCACAGUGCUCCCAACCGCUCUGUUUGCCUUGGCAAUUAGCCGGAAUUCCCCGCACUUU-GCAACCGCAGAUA .............(((((((((.......((((.((........)).))))((((....))))...)))))........((.....-))..))))..... ( -26.60) >DroSim_CAF1 6247 99 + 1 UGUUCAAUCAUAAGCGGCUGGCCAUAAUCACAGUGCUCCCAACCGCUCUGUUUGCCUUGGCAAUUAGCCGGAAUUCCCCGCACUUU-GCAACCGCAGAUA .............(((((((((.......((((.((........)).))))((((....))))...)))))........((.....-))..))))..... ( -26.60) >DroEre_CAF1 6208 99 + 1 UGUUCAAUCAUAAGCGGCUGGCCAUAAUCACAGUGCUCCCAACCGCUCUGUUUGCCUUGGCAAUUAGCCAGAAUUCCCCGCACAUU-GCCACCGCAGAUA ............(((((.(((.(((.......)))...))).))))).(((((((..(((((((..((..(.....)..))..)))-))))..))))))) ( -26.50) >consensus UGUUCAAUCAUAAGCGGCUGGCCAUAAUCACAGUGCUCCCAACCGCUCUGUUUGCCUUGGCAAUUAGCCAGAAUUCCCCGCACUUU_GCAACCGCAGAUA ......(((....(((((((((.......((((.((........)).))))((((....))))...)))))......))))......((....)).))). (-25.44 = -25.00 + -0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:16 2006