
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,902,160 – 13,902,301 |
| Length | 141 |
| Max. P | 0.590023 |

| Location | 13,902,160 – 13,902,261 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 87.41 |
| Mean single sequence MFE | -24.23 |
| Consensus MFE | -21.84 |
| Energy contribution | -22.53 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.590023 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
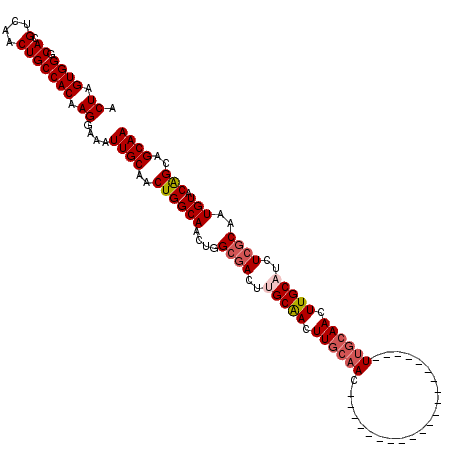
>X_DroMel_CAF1 13902160 101 + 22224390 UUCCAAGGUGCGAUACCCCAAACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUUGCAACUUGCAACUAGCAACUCGUAAC ...((((.(((((...((((......))))..(((((..((((((((.....)))....)))))..)))))..))))).)))).................. ( -25.70) >DroSec_CAF1 2774 87 + 1 UUCCAAGGUGCGAUACCCCAAACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUUGCAACUUGCAAC-------------- ...((((.(((((...((((......))))..(((((..((((((((.....)))....)))))..)))))..))))).))))....-------------- ( -25.70) >DroSim_CAF1 24538 82 + 1 UUCCAAGGUGCGAUACCCCAAACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUCGCAACUU------------------- ....(((.(((((...((((......))))..(((((..((((((((.....)))....)))))..)))))..))))).)))------------------- ( -24.80) >DroYak_CAF1 12756 87 + 1 UUCCAAGGUGCGAUACCCCAAACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUUGCAACUGGCGACUUGCAAC-------------- .....((.((((((..((((......))((((.(....)))))....))..)))))).)).((((.((((.....)))).))))...-------------- ( -20.70) >consensus UUCCAAGGUGCGAUACCCCAAACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUUGCAACUUGCAAC______________ ...((((.(((((...((((......))))..(((((..((((((((.....)))....)))))..)))))..))))).)))).................. (-21.84 = -22.53 + 0.69)



| Location | 13,902,181 – 13,902,301 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.50 |
| Mean single sequence MFE | -28.55 |
| Consensus MFE | -22.29 |
| Energy contribution | -23.73 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.78 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.512256 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13902181 120 + 22224390 ACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUUGCAACUUGCAACUAGCAACUCGUAACUCGCAACUCGCAACUUGCAUCUCGCAAUGUGCAGCAGCAA ....((..((((((((..((((((((.....)))....)))))..))((((..(((((.((((.....((............)).....)))).)))))..)))).)))))).))..... ( -32.70) >DroSec_CAF1 2795 99 + 1 ACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUUGCAACUUGCAAC---------------------UUGCAACUUGCAUUUCGCAAUGUACGGCAGCAA .((.((((.((.(....))))))).))....((((..((((((....((((..(((((.((((...---------------------..)))).)))))..))))..))).)))..)))) ( -31.20) >DroSim_CAF1 24559 92 + 1 ACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUCGCAACUU----------------------------GCAACUUGCAUCUCGCAAUGUACGGCAGCAA ....(((((..(((((..((((((((.....)))....)))))..))))).)))))...((----------------------------((.((((((.....)))).))...))))... ( -25.30) >DroYak_CAF1 12777 99 + 1 ACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUUGCAACUGGCGACUUGCAAC---------------------UUGCAACUUGCAUCUCACAAUGUUCAGCAGCAA ....(((((((.(....)))))....(.((.((((((...((((.((((.....)))).))))...---------------------)))))).)).)....)))..(((......))). ( -25.00) >consensus ACUAGUGGGCACGUCAACUGCCACAAGGAAAUUGCAACUGGCAACUGGCGACUUGCAACUUGCAAC_____________________UUGCAACUUGCAUCUCGCAAUGUACAGCAGCAA .((.((((.((.(....))))))).))....((((..((((((....((((..(((((.((((((......................)))))).)))))..))))..))).)))..)))) (-22.29 = -23.73 + 1.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:40 2006