| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,518,099 – 1,518,200 |
| Length | 101 |
| Max. P | 0.998935 |

| Location | 1,518,099 – 1,518,200 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.91 |
| Mean single sequence MFE | -36.80 |
| Consensus MFE | -27.58 |
| Energy contribution | -28.30 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.75 |
| SVM decision value | 3.29 |
| SVM RNA-class probability | 0.998935 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
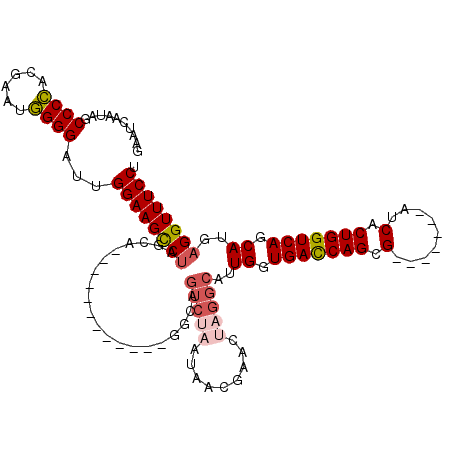
>X_DroMel_CAF1 1518099 101 + 22224390 GAAUCAAUAGCCCCACGAACGGGGAUUGGAAGCCUAGCA------------GGCAGUCUAAUAACGAACUAGGCAUUGGUGACCAGCG-------AUCACUGGUCAGCAUGAGGUUUCCU ..........((((......))))...((((((((....------------.((.(((((.........))))).....(((((((..-------....)))))))))...)))))))). ( -36.30) >DroSec_CAF1 7548 101 + 1 GAAUCAAUAGCCCCACGAAUGGGGAUUGGAAGCCUAGCA------------GGCAGUCUAAUAACUAACUAGGCAUUGGUGACCAGCG-------AUCACUGGUCAGCAUGAGGUUUCCU ..........(((((....)))))...((((((((....------------.((.(((((.........))))).....(((((((..-------....)))))))))...)))))))). ( -36.80) >DroSim_CAF1 8811 101 + 1 GAAUCAAUAGCCCCACGAAUGGGGAUUGGAAGCCUAGCA------------GGCAGUCUAAUAACUAACUAGGCAUUGGUGACCAGCG-------AUCACUGGUCAGCAUGAGGUUUCCU ..........(((((....)))))...((((((((....------------.((.(((((.........))))).....(((((((..-------....)))))))))...)))))))). ( -36.80) >DroEre_CAF1 7674 113 + 1 GAAACAAUAGCCCUACGAAUAGGGAUUGGAAGCCCGGGAUCGAAAGCCCCGUGCAGUC-------GAAAUAGGCAUUGGUGAUCAGCGGUCGGCGAUCACUGGUCACCAUGAGGUUUCCU ..........(((((....)))))...(((((((((((..(....).))))....(((-------......)))..((((((((((.((((...)))).))))))))))...))))))). ( -45.50) >DroYak_CAF1 9237 88 + 1 GAAACAAUAGCCCUAUGACUAGGGAUUGGAAGUCU--------------------------------AAUAGGCAUUGGUGAUCAGCGAUCAGCUAUCACUGGUCAGCAUGAGGUUUCCU ..........(((((....)))))...((((..((--------------------------------.....((((..(((((.(((.....))))))))..))..))...))..)))). ( -28.60) >consensus GAAUCAAUAGCCCCACGAAUGGGGAUUGGAAGCCUAGCA____________GGCAGUCUAAUAACGAACUAGGCAUUGGUGACCAGCG_______AUCACUGGUCAGCAUGAGGUUUCCU ..........((((......))))...((((((((....................(((((.........)))))..((.(((((((.(.........).))))))).))..)))))))). (-27.58 = -28.30 + 0.72)

| Location | 1,518,099 – 1,518,200 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.91 |
| Mean single sequence MFE | -34.60 |
| Consensus MFE | -27.19 |
| Energy contribution | -28.19 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.79 |
| SVM decision value | 3.25 |
| SVM RNA-class probability | 0.998852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
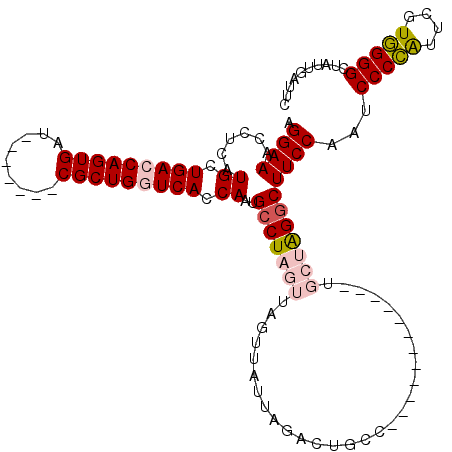
>X_DroMel_CAF1 1518099 101 - 22224390 AGGAAACCUCAUGCUGACCAGUGAU-------CGCUGGUCACCAAUGCCUAGUUCGUUAUUAGACUGCC------------UGCUAGGCUUCCAAUCCCCGUUCGUGGGGCUAUUGAUUC .((((......((.((((((((...-------.)))))))).))..(((((((..((.........)).------------.)))))))))))...(((((....))))).......... ( -36.20) >DroSec_CAF1 7548 101 - 1 AGGAAACCUCAUGCUGACCAGUGAU-------CGCUGGUCACCAAUGCCUAGUUAGUUAUUAGACUGCC------------UGCUAGGCUUCCAAUCCCCAUUCGUGGGGCUAUUGAUUC .((((......((.((((((((...-------.)))))))).))..((((((((((((....)))))..------------.)))))))))))...(((((....))))).......... ( -38.90) >DroSim_CAF1 8811 101 - 1 AGGAAACCUCAUGCUGACCAGUGAU-------CGCUGGUCACCAAUGCCUAGUUAGUUAUUAGACUGCC------------UGCUAGGCUUCCAAUCCCCAUUCGUGGGGCUAUUGAUUC .((((......((.((((((((...-------.)))))))).))..((((((((((((....)))))..------------.)))))))))))...(((((....))))).......... ( -38.90) >DroEre_CAF1 7674 113 - 1 AGGAAACCUCAUGGUGACCAGUGAUCGCCGACCGCUGAUCACCAAUGCCUAUUUC-------GACUGCACGGGGCUUUCGAUCCCGGGCUUCCAAUCCCUAUUCGUAGGGCUAUUGUUUC .((((......((((((.(((((.((...)).))))).))))))..((((...((-------((..((.....))..))))....))))))))...(((((....))))).......... ( -37.20) >DroYak_CAF1 9237 88 - 1 AGGAAACCUCAUGCUGACCAGUGAUAGCUGAUCGCUGAUCACCAAUGCCUAUU--------------------------------AGACUUCCAAUCCCUAGUCAUAGGGCUAUUGUUUC ..(((((....((.(((.(((((((.....))))))).))).))..((((...--------------------------------.((((..........))))...))))....))))) ( -21.80) >consensus AGGAAACCUCAUGCUGACCAGUGAU_______CGCUGGUCACCAAUGCCUAGUUAGUUAUUAGACUGCC____________UGCUAGGCUUCCAAUCCCCAUUCGUGGGGCUAUUGAUUC .((((......((.(((((((((.........))))))))).))..(((((((.............................)))))))))))...(((((....))))).......... (-27.19 = -28.19 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:47:03 2006