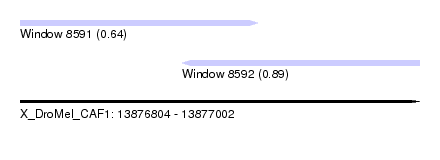
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,876,804 – 13,877,002 |
| Length | 198 |
| Max. P | 0.891923 |

| Location | 13,876,804 – 13,876,922 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.48 |
| Mean single sequence MFE | -27.33 |
| Consensus MFE | -20.78 |
| Energy contribution | -20.67 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
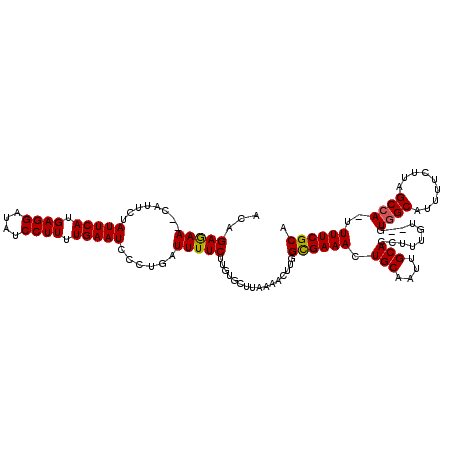
>X_DroMel_CAF1 13876804 118 + 22224390 ACAGAGAAUUCAUUUUAUUCAUGAGGAAAUCCUUUUGAAUUCCUUAUUUUCUUGUGCUUAAAACUUGGUGAAACUGCAAUUGCACCUUUCU--GUUGCGUUUUCUUAGCCAAUUUUCGCA (((((((((.......(((((.((((....)))).))))).....)))))).)))((..((((.(((((((((.((((((...........--)))))).))))...))))))))).)). ( -26.50) >DroSec_CAF1 26097 116 + 1 ACAGAGAA--CAUUCUAUUCACGAGGAUAUCCUUUUGAAUCCCUGAUUUUCUUGUGCUUAAAACUUGGCGAAACUGCAAUUGCACCUUUGU--GUGGCAUUUUCUUAGCCAUUUUUCGCA ((((((((--((....(((((.((((....)))).)))))...)).))))).)))((..((((..((((((((.(((....((((....))--)).))).))))...))))))))..)). ( -27.90) >DroEre_CAF1 20901 116 + 1 A-AGAAAA--CAUUCUAUUCAUGAGGAUAUCCUUUUGAAUCCCUGAUUUUCCUGUGCUGCAAACUUGGCGAAACUGCAAUUGCACCUUUGUGUGUGGCAUUUUCUUAGCCA-UUUUCGCU .-.(((((--((....(((((.((((....)))).)))))...)).))))).......((.((..((((((((.(((...(((((....)))))..))).))))...))))-..)).)). ( -27.60) >consensus ACAGAGAA__CAUUCUAUUCAUGAGGAUAUCCUUUUGAAUCCCUGAUUUUCUUGUGCUUAAAACUUGGCGAAACUGCAAUUGCACCUUUGU__GUGGCAUUUUCUUAGCCA_UUUUCGCA ...(((((........(((((.((((....)))).)))))......)))))................((((((.(((....)))..........((((.........))))..)))))). (-20.78 = -20.67 + -0.11)

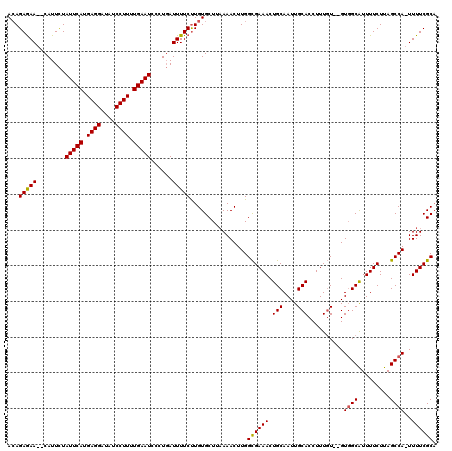
| Location | 13,876,884 – 13,877,002 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.00 |
| Mean single sequence MFE | -29.70 |
| Consensus MFE | -24.73 |
| Energy contribution | -26.40 |
| Covariance contribution | 1.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.94 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.891923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
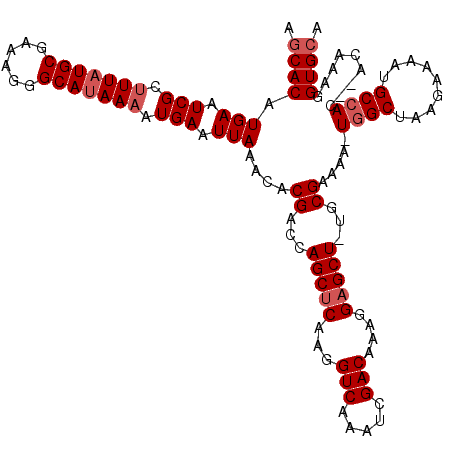
>X_DroMel_CAF1 13876884 118 - 22224390 AGCACAUGAAUCGCUUUAUGCGAAAGGGCAUAAAAUGAAUUAAACACGACCAGCUCAAGGUCAAAUCGACAAAGGCGCUUUGCGAAAAUUGGCUAAGAAAACGCAAC--AGAAAGGUGCA .((((.(((.(((.(((((((......))))))).))).)))..........((((..((((((.(((.(((((...))))))))...))))))..))....))...--......)))). ( -27.20) >DroSec_CAF1 26175 117 - 1 AGCACAUGAAUCGCUUUAUGCGAAAGGGCAUAAAAUGAAUUAAACACGACCAGCUCAAGGUCAAAUCGACAAAGGAGCU-UGCGAAAAAUGGCUAAGAAAAUGCCAC--ACAAAGGUGCA .((((.....(((((((((((......))))))).................(((((...(((.....)))....)))))-.))))....((((.........)))).--......)))). ( -33.00) >DroEre_CAF1 20978 118 - 1 AACACAUGAAUCGCUUUAUGCGAAAGGGCACAACAUGAAUUAAACACGACCAGCUCAAGGUCAAAUCGACAAAGGAGCU-AGCGAAAA-UGGCUAAGAAAAUGCCACACACAAAGGUGCA ..........((((.....))))....((((...............((.(.(((((...(((.....)))....)))))-.)))....-((((.........)))).........)))). ( -28.90) >consensus AGCACAUGAAUCGCUUUAUGCGAAAGGGCAUAAAAUGAAUUAAACACGACCAGCUCAAGGUCAAAUCGACAAAGGAGCU_UGCGAAAA_UGGCUAAGAAAAUGCCAC__ACAAAGGUGCA .((((.(((.(((.(((((((......))))))).))).)))....((...(((((...(((.....)))....)))))...)).....((((.........)))).........)))). (-24.73 = -26.40 + 1.67)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:32 2006