
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,704,254 – 13,704,374 |
| Length | 120 |
| Max. P | 0.783382 |

| Location | 13,704,254 – 13,704,374 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.11 |
| Mean single sequence MFE | -33.00 |
| Consensus MFE | -31.52 |
| Energy contribution | -31.55 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.03 |
| Mean z-score | -0.89 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.710830 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13704254 120 + 22224390 ACCGGUGUGCACGCGACAAUGAUCUUUCAUGUGGUGGCGCAGCUUAAAUGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU ........(((((((((.((((....))))((.((((((.(((......))).))))))...)).............)))((((((.(((.(((....))).))).)))))).))).))) ( -34.00) >DroPse_CAF1 18172 120 + 1 ACCGGUGUGGACGCGACAAUGAUCUUUCAUGUGGUGGCGCAGCUUAAACGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU ((((((((((.(((....((((....)))))))((((((.(((......))).)))))).....)))))).....)))).((((((.(((.(((....))).))).))))))........ ( -33.00) >DroGri_CAF1 21618 120 + 1 ACCCGUGUGCACGCGACAAUGAUCUUUCAUGUGGUGGCGCAGCUUAAACGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU ........(((((((((.((((....))))((.((((((.(((......))).))))))...)).............)))((((((.(((.(((....))).))).)))))).))).))) ( -33.40) >DroWil_CAF1 21619 120 + 1 ACCUGUGUGCACGCGACAAUGGUCUUUCAUGUGAUGGCGCAGCUUAAACGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU .((((....(((((((((.(((((..(((((((((((((.(((......))).)))))......)))))......)))..)))))..))).(((....)))..))))))..))).).... ( -31.20) >DroMoj_CAF1 20348 120 + 1 ACCCGUGUGCACGCGACAAUGAUCUUUCAUGUGGUGGCGCAGCUUAAACGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU ........(((((((((.((((....))))((.((((((.(((......))).))))))...)).............)))((((((.(((.(((....))).))).)))))).))).))) ( -33.40) >DroPer_CAF1 18106 120 + 1 ACCGGUGUGGACGCGACAAUGAUCUUUCAUGUGGUGGCGCAGCUUAAACGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU ((((((((((.(((....((((....)))))))((((((.(((......))).)))))).....)))))).....)))).((((((.(((.(((....))).))).))))))........ ( -33.00) >consensus ACCGGUGUGCACGCGACAAUGAUCUUUCAUGUGGUGGCGCAGCUUAAACGCUUUGCCACAAAACUCACAUUUAAAUGGUCGAUCACCUGUAUGUACCGACAAAUGCGUGAUCAGUGAUGU ...........((((((.((((....))))((.((((((.(((......))).))))))...)).............)))((((((.(((.(((....))).))).)))))).))).... (-31.52 = -31.55 + 0.03)
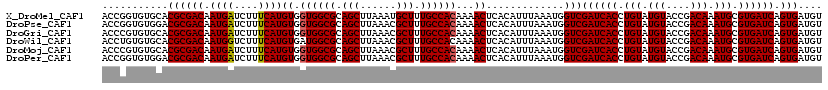


| Location | 13,704,254 – 13,704,374 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.11 |
| Mean single sequence MFE | -33.70 |
| Consensus MFE | -31.37 |
| Energy contribution | -31.40 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.783382 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13704254 120 - 22224390 ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCAUUUAAGCUGCGCCACCACAUGAAAGAUCAUUGUCGCGUGCACACCGGU ....((.(((((.(((((((((((((.(((....))))))))).....)))))))......(((((..(((......)))..))))).........))))).))(((.((....))))). ( -33.00) >DroPse_CAF1 18172 120 - 1 ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCGUUUAAGCUGCGCCACCACAUGAAAGAUCAUUGUCGCGUCCACACCGGU ....((.(((((.(((((((((((((.(((....))))))))).....)))))))......(((((..(((......)))..))))).........))))).))(((.((....))))). ( -32.60) >DroGri_CAF1 21618 120 - 1 ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCGUUUAAGCUGCGCCACCACAUGAAAGAUCAUUGUCGCGUGCACACGGGU ....((.(((((.(((((((((((((.(((....))))))))).....)))))))......(((((..(((......)))..))))).........))))).)).(.(((....))).). ( -34.80) >DroWil_CAF1 21619 120 - 1 ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCGUUUAAGCUGCGCCAUCACAUGAAAGACCAUUGUCGCGUGCACACAGGU ......(((..(((((....((((((.(((....))))))))).......((((((......((((..(((......)))..))))))))))....(((....))))))))....))).. ( -34.40) >DroMoj_CAF1 20348 120 - 1 ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCGUUUAAGCUGCGCCACCACAUGAAAGAUCAUUGUCGCGUGCACACGGGU ....((.(((((.(((((((((((((.(((....))))))))).....)))))))......(((((..(((......)))..))))).........))))).)).(.(((....))).). ( -34.80) >DroPer_CAF1 18106 120 - 1 ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCGUUUAAGCUGCGCCACCACAUGAAAGAUCAUUGUCGCGUCCACACCGGU ....((.(((((.(((((((((((((.(((....))))))))).....)))))))......(((((..(((......)))..))))).........))))).))(((.((....))))). ( -32.60) >consensus ACAUCACUGAUCACGCAUUUGUCGGUACAUACAGGUGAUCGACCAUUUAAAUGUGAGUUUUGUGGCAAAGCGUUUAAGCUGCGCCACCACAUGAAAGAUCAUUGUCGCGUGCACACCGGU .(..((.(((((.(((((((((((((.(((....))))))))).....)))))))......(((((..(((......)))..))))).........))))).))..)............. (-31.37 = -31.40 + 0.03)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:43:18 2006