
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,628,157 – 13,628,383 |
| Length | 226 |
| Max. P | 0.996645 |

| Location | 13,628,157 – 13,628,276 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.72 |
| Mean single sequence MFE | -23.20 |
| Consensus MFE | -20.21 |
| Energy contribution | -20.69 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.73 |
| SVM RNA-class probability | 0.996645 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13628157 119 - 22224390 UUUCCCCUCGUCGACGUUUAUGAAUAACUUUAUAAAACCAGACCGCACACUCUGAUUGCGGUUUCCACUUCCUUUUCUGCCAUAUUAUAUAGUUUCAGCAUUCGCGUGACGCACUCCCU- ........((((.((((..(((...((((.(((((....((((((((.........)))))))).......(......).....))))).))))....)))..))))))))........- ( -23.20) >DroSec_CAF1 21797 119 - 1 UUUCCCCUCGUCACCGUUUAUGAAUAACUUUAUAAAACCAGACCGCACACUCUGAUUGUGGUUUCCACUUCCUUUUCUGCCAUAUUUUAUAGUUUAAGCAUUCGCGUGACGCACACCCU- ........((((((((...(((..(((..((((((((.((((........))))..((((((................)))))))))))))).)))..))).)).))))))........- ( -23.29) >DroSim_CAF1 15043 119 - 1 UUUCCCCUCGUCACCGUUUAUGAAUAACUUUAUAAAACCAGACCGCACACUCUGAUUGUGGUUUCCACUUCCUUUUCUGCCAUAUUAUAUAGUUUCAGCAUUCGCGUGACGCACUCCCU- ........((((((((...(((...((((.(((((...((((........))))..((((((................))))))))))).))))....))).)).))))))........- ( -24.59) >DroEre_CAF1 13179 120 - 1 UUUCCCCUCGCCAUCGUUUAUGAAUAACUUUAUAAACCCAGACCGCACACUCUGAUUGUGGUUUCCACUUCCGUUUCUGCCAUAUUAUAUAGUUUCAGCAUUCGCGUGACGCAUUCCCUC .........((...(((..(((...((((.(((((...((((........))))..((((((....((....))....))))))))))).))))....)))..)))....))........ ( -20.50) >DroYak_CAF1 12490 118 - 1 -UUCCCCUCGUCACCGUUUAUGAAUAACUUUAUAAACCCUGGCCGCACACUCUGAUUGUGGUUUCCACUUCCGUUUCUGCCAUAUUAUAUAGUUUCAGCAUUCGCGUGACGCACUCCCU- -.......((((((((...(((...((((.(((((....((((.(...((...((..((((...)))).)).))..).))))..))))).))))....))).)).))))))........- ( -24.40) >consensus UUUCCCCUCGUCACCGUUUAUGAAUAACUUUAUAAAACCAGACCGCACACUCUGAUUGUGGUUUCCACUUCCUUUUCUGCCAUAUUAUAUAGUUUCAGCAUUCGCGUGACGCACUCCCU_ ........(((((.(((..(((...((((.(((((...((((........))))..((((((................))))))))))).))))....)))..))))))))......... (-20.21 = -20.69 + 0.48)



| Location | 13,628,276 – 13,628,383 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.95 |
| Mean single sequence MFE | -22.28 |
| Consensus MFE | -14.44 |
| Energy contribution | -14.28 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.737886 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13628276 107 - 22224390 CCACGGAAACUGCAGUUUGCCCUCU-GGGGUUUAAUUGCACUGC---ACCCGCUUCCGUCCAUUUUGCAUUUUACUCUUUACCAUCU---------ACCUACCAUCUAUUCAGCGCCUCU ..((((((..(((((((.((((...-.))))..)))))))..((---....))))))))............................---------........................ ( -20.60) >DroSec_CAF1 21916 106 - 1 CCACGGAAACUGCAGUUUGCCCUGC-GGGGGUUAAUUGCACUGC---CCCCGCUUCCGCCCAUUUUGCAUUUUACUCUUUACCUCCU---------AACUACCAUCUAUGCAGCGCC-CU ....((...(((((.........((-((((((..........))---))))))....((.......))...................---------............)))))..))-.. ( -26.20) >DroSim_CAF1 15162 115 - 1 CCACGGAAACUGCAGUUUGCCCUCC-GGGGGUUAAUUGCACUGC---CCCCGCUUCCGCCCAUUUUGCAUUUUACUCUUUACCUCCCUUUACCUCCUCCUACCAUCUAUGCAGCGCC-CU ....((...(((((..........(-((((((..........))---))))).....((.......))........................................)))))..))-.. ( -22.30) >DroEre_CAF1 13299 100 - 1 CCGCGGAAACUGCAGUUUGACCUCC-GGGGGUUAAUUGCACUGC--CUCCCGUUUCCGUCCAUUUUGCAUUUUACUCUUUACCUUCU---------ACCAUCCAGU-------CGCC-CU ..((((((((.((((((((((((..-..)))))))....)))))--.....))))))))............................---------..........-------....-.. ( -22.60) >DroYak_CAF1 12608 97 - 1 CCACGGAAACUGCAGUUCGGCCUCCAGGGGGUUAAUUGCACCGCACCCCCCGCUUCCGUUCAUUUUGCAUUUUACUCUUUACCCUCU---------ACCAACCAGC-------------- ..((((((..(((((((.(((((.....))))))))))))..((.......))))))))............................---------..........-------------- ( -19.70) >consensus CCACGGAAACUGCAGUUUGCCCUCC_GGGGGUUAAUUGCACUGC___CCCCGCUUCCGUCCAUUUUGCAUUUUACUCUUUACCUUCU_________ACCUACCAUCUAU_CAGCGCC_CU ...(((((..(((((((...(((....)))...)))))))..((.......))))))).............................................................. (-14.44 = -14.28 + -0.16)


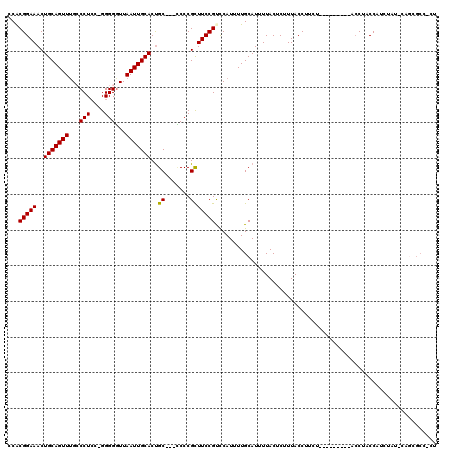
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:40 2006