
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,616,345 – 13,616,481 |
| Length | 136 |
| Max. P | 0.983285 |

| Location | 13,616,345 – 13,616,441 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 92.81 |
| Mean single sequence MFE | -28.97 |
| Consensus MFE | -24.04 |
| Energy contribution | -24.20 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.582704 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13616345 96 + 22224390 CGGUAGGCCAUCGCAGAUAACAGAGCUCUGUUAGCUGUGCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGG .((....))..((((.......(((((((..((((((.(((..(((((....))))))))))))))...........))))))).......)))). ( -33.06) >DroSec_CAF1 2217 96 + 1 CGGUAGGCCAUCGCAGAUAACAGAGCUCUGUUAACAGACCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGG .((....))..((((.......(((((((.(((.((((..((.(((.(((((.....)))))))).)).)))).)))))))))).......)))). ( -29.54) >DroSim_CAF1 2205 96 + 1 CGGCAGGCCAUCGCAGAUAACAGAGCUCUGUUAACAGACCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAACUCGCAAAGAUGCGG .(((.(((((..((...((((((....)))))).((.(((...))).)))).....))))).)))..................((((....)))). ( -25.70) >DroEre_CAF1 2175 95 + 1 CGAUAGGCCA-CGCAGAUAACAGAGCUCUGUUAACAGAGCCAAGGUGUGGCUACACUGGCAAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGG ..........-((((.......((((((((((.....)))((((((((((((.........))))))))))))....))))))).......)))). ( -29.64) >DroYak_CAF1 2119 95 + 1 CGAUAAGCCAUCGCAG-UAACAGAGCUCUGUUAACAAAGCCAUGGUGUGGCGACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGG ...........((((.-.....(((((((.(((.(((((..(((((.((((.......))))))))).))))).)))))))))).......)))). ( -26.92) >consensus CGGUAGGCCAUCGCAGAUAACAGAGCUCUGUUAACAGAGCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGG ...........((((.......((((((((....).....(((((((((((...........)))))))))))....))))))).......)))). (-24.04 = -24.20 + 0.16)

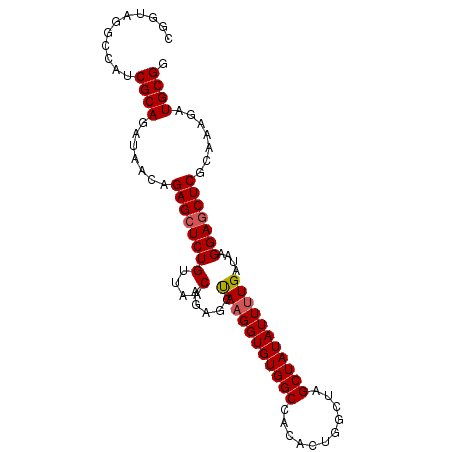

| Location | 13,616,345 – 13,616,441 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 92.81 |
| Mean single sequence MFE | -26.28 |
| Consensus MFE | -21.90 |
| Energy contribution | -22.22 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.707956 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13616345 96 - 22224390 CCGCAUCUUUGCGAGCUCCUUAUCAAAAUAUAGCUAGCCAGUGUGGCCACACCUUAGCACAGCUAACAGAGCUCUGUUAUCUGCGAUGGCCUACCG .((((....))))((((...(((....))).)))).((((((((....)))).....(.(((.((((((....)))))).))).).))))...... ( -25.40) >DroSec_CAF1 2217 96 - 1 CCGCAUCUUUGCGAGCUCCUUAUCAAAAUAUAGCUAGCCAGUGUGGCCACACCUUAGGUCUGUUAACAGAGCUCUGUUAUCUGCGAUGGCCUACCG .((((....))))((((...(((....))).)))).((((.(((((((........))))...((((((....))))))...))).))))...... ( -24.20) >DroSim_CAF1 2205 96 - 1 CCGCAUCUUUGCGAGUUCCUUAUCAAAAUAUAGCUAGCCAGUGUGGCCACACCUUAGGUCUGUUAACAGAGCUCUGUUAUCUGCGAUGGCCUGCCG .((((....))))...................((..((((.(((((((........))))...((((((....))))))...))).))))..)).. ( -25.50) >DroEre_CAF1 2175 95 - 1 CCGCAUCUUUGCGAGCUCCUUAUCAAAAUAUAGCUUGCCAGUGUAGCCACACCUUGGCUCUGUUAACAGAGCUCUGUUAUCUGCG-UGGCCUAUCG ..((((....(((((((...(((....))).)))))))..)))).(((((.....(((((((....)))))))..((.....)))-))))...... ( -29.20) >DroYak_CAF1 2119 95 - 1 CCGCAUCUUUGCGAGCUCCUUAUCAAAAUAUAGCUAGCCAGUGUCGCCACACCAUGGCUUUGUUAACAGAGCUCUGUUA-CUGCGAUGGCUUAUCG .((((....))))((((...(((....))).))))(((((...((((.....((.(((((((....))))))).))...-..)))))))))..... ( -27.10) >consensus CCGCAUCUUUGCGAGCUCCUUAUCAAAAUAUAGCUAGCCAGUGUGGCCACACCUUAGCUCUGUUAACAGAGCUCUGUUAUCUGCGAUGGCCUACCG .((((....))))((((...(((....))).)))).((((.(((((..(((....(((((((....))))))).)))...))))).))))...... (-21.90 = -22.22 + 0.32)

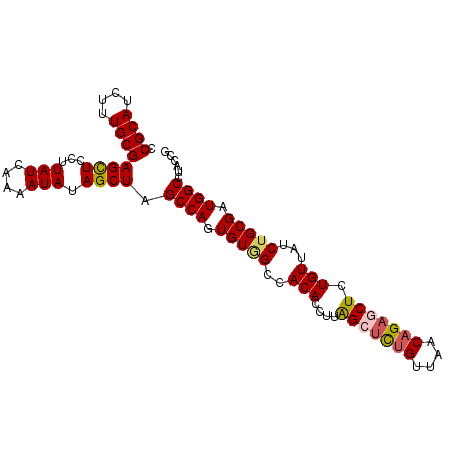
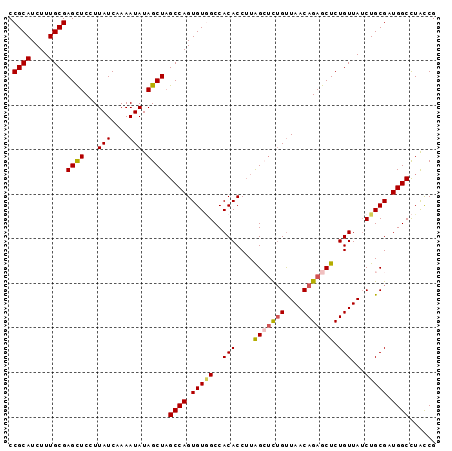
| Location | 13,616,361 – 13,616,481 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.83 |
| Mean single sequence MFE | -39.98 |
| Consensus MFE | -35.00 |
| Energy contribution | -35.52 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983285 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13616361 120 + 22224390 AUAACAGAGCUCUGUUAGCUGUGCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGGAUUUUCUACGGACUACUGGCGUUCCUGGUGGCCAGACAAC .((((((....))))))...........((.(((((((...((...((((((.((((.((..(((..((((....))))..)))..)))))).)).))))...))..))))))).))... ( -40.20) >DroSec_CAF1 2233 120 + 1 AUAACAGAGCUCUGUUAACAGACCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGGAUUUUCUACGGAUUGCUGGCGUUCCUGGUGGCCAGACAGC .((((((....))))))...........((.(((((((....((((((.....((((.((..(((..((((....))))..)))..))))))..)))))).......))))))).))... ( -40.90) >DroSim_CAF1 2221 120 + 1 AUAACAGAGCUCUGUUAACAGACCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAACUCGCAAAGAUGCGGAUUUUCUACGGAUUGCUGGCGUUCCUGGUGGCCAGACAGC .((((((....))))))...........((.(((((((....((((((.....((((.((..(((..((((....))))..)))..))))))..)))))).......))))))).))... ( -41.20) >DroEre_CAF1 2190 120 + 1 AUAACAGAGCUCUGUUAACAGAGCCAAGGUGUGGCUACACUGGCAAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGGAUUUUCUACGGAUUUCUGGCGUUCCUGAUGGCCAGCCAAC ........((((((....))))))((((((((((((.........))))))))))))...((((((.((((....))))((...((....))..))....))))))..(((....))).. ( -38.90) >DroYak_CAF1 2135 119 + 1 -UAACAGAGCUCUGUUAACAAAGCCAUGGUGUGGCGACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGGAUUUUCUACGGAAUUCUGGCGUUCCUGGUGGCCAGCCGAC -((((((....)))))).....(((((...))))).....(((((.(((((.........((((((.((((....))))((..(((....))).))....)))))).))))).))))).. ( -38.70) >consensus AUAACAGAGCUCUGUUAACAGAGCUAAGGUGUGGCCACACUGGCUAGCUAUAUUUUGAUAAGGAGCUCGCAAAGAUGCGGAUUUUCUACGGAUUGCUGGCGUUCCUGGUGGCCAGACAAC .((((((....))))))...........((.(((((((...((...((((...((((.((..(((..((((....))))..)))..))))))....))))...))..))))))).))... (-35.00 = -35.52 + 0.52)
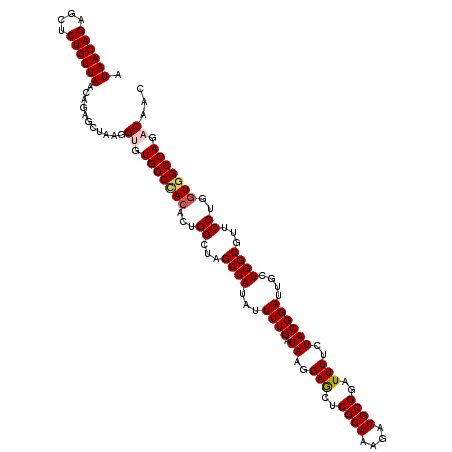


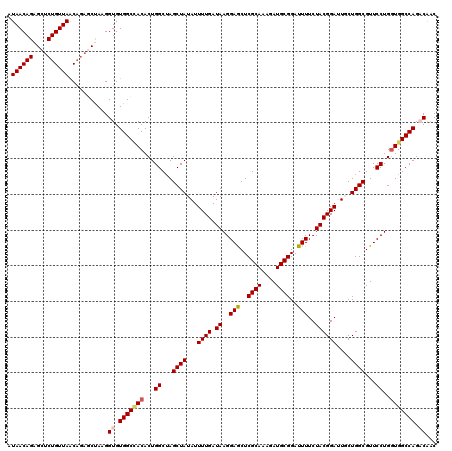
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:16 2006