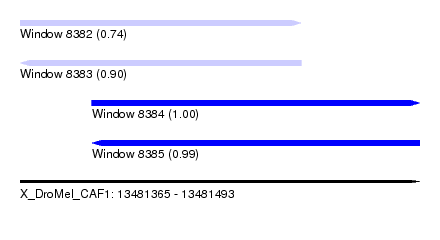

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,481,365 – 13,481,493 |
| Length | 128 |
| Max. P | 0.999999 |

| Location | 13,481,365 – 13,481,455 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 86.46 |
| Mean single sequence MFE | -33.13 |
| Consensus MFE | -27.67 |
| Energy contribution | -28.33 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.742004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13481365 90 + 22224390 CGGAUCCCAACCCGUGUAUCUGCCUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGCUUUUGUG------ ((((((((.(((.(.((((((((.......))))..))))).)))...(((.....)))))))))))....((((....)))).......------ ( -36.80) >DroSec_CAF1 14147 96 + 1 CCGAUCCCAAACCGUGUAUCUGCCUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGCUUAUGUGGCUCUA ..........(((..((((((((.......))))..))))..)))..(((((((..(((((((((((.....)))).))))))).))).))))... ( -35.80) >DroSim_CAF1 6757 96 + 1 CCGAUCCCAAACCGUGUAUCUGCCUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGNNNNNNNNNNNNNN ..((((((..(((..((((((((.......))))..))))..)))...(((.....))))))))).......(((....))).............. ( -26.80) >consensus CCGAUCCCAAACCGUGUAUCUGCCUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGCUU_UGUG______ ..((((((..(((..((((((((.......))))..))))..)))...(((.....)))))))))......((((....))))............. (-27.67 = -28.33 + 0.67)

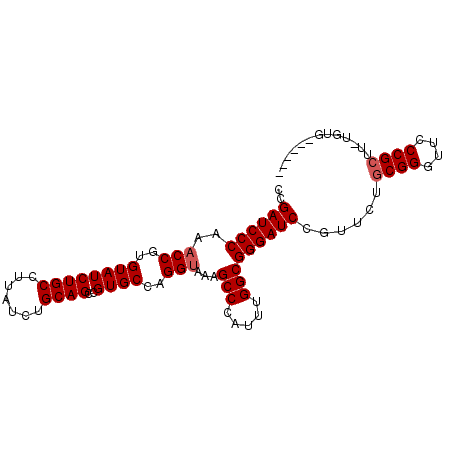
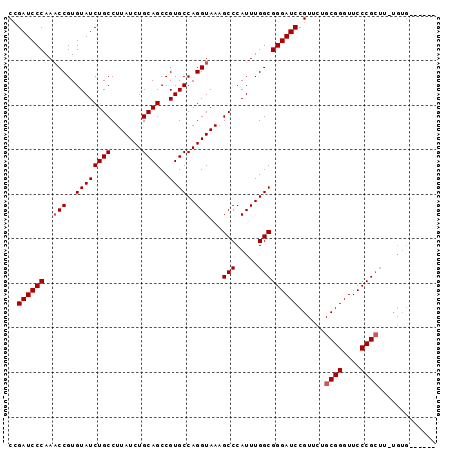
| Location | 13,481,365 – 13,481,455 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 86.46 |
| Mean single sequence MFE | -34.30 |
| Consensus MFE | -29.93 |
| Energy contribution | -30.27 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.897292 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13481365 90 - 22224390 ------CACAAAAGCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAGGCAGAUACACGGGUUGGGAUCCG ------.......((((....))))....(((((((((((.....)))...(((((......((((.......)))).....))))).)))))))) ( -39.10) >DroSec_CAF1 14147 96 - 1 UAGAGCCACAUAAGCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAGGCAGAUACACGGUUUGGGAUCGG (((((((.(((..((((((..(((.....))).))))))...))))))))))((..(..((.((((.......)))).....))..)..))..... ( -36.00) >DroSim_CAF1 6757 96 - 1 NNNNNNNNNNNNNNCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAGGCAGAUACACGGUUUGGGAUCGG ..............(((....))).....(.((((((((((...((......)))))).((.((((.......)))).....))....)))))).) ( -27.80) >consensus ______CACA_AAGCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAGGCAGAUACACGGUUUGGGAUCGG .............((((....))))......((((((((((...((......)))))).((.((((.......)))).....))....)))))).. (-29.93 = -30.27 + 0.33)

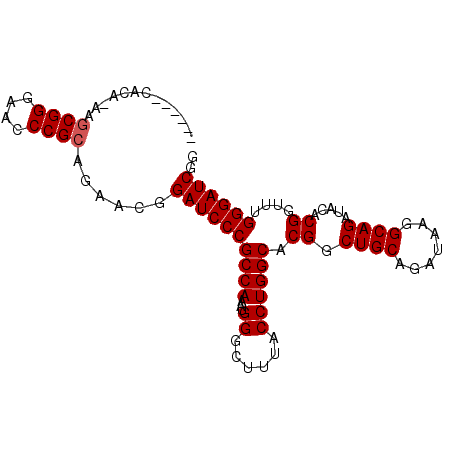
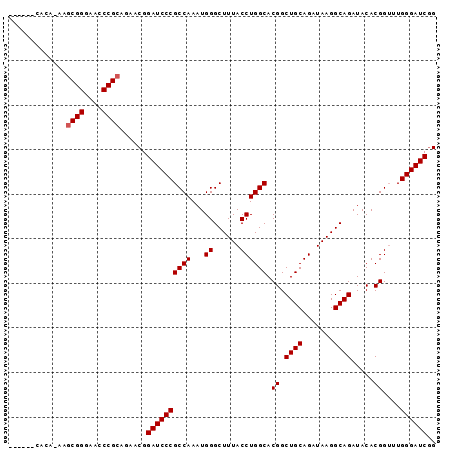
| Location | 13,481,388 – 13,481,493 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 65.09 |
| Mean single sequence MFE | -39.47 |
| Consensus MFE | -24.95 |
| Energy contribution | -26.20 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.63 |
| SVM decision value | 6.68 |
| SVM RNA-class probability | 0.999999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13481388 105 + 22224390 CUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGCUUUUGUG--------UGCAGGUAGUUGGGCGCGUUCCGCAAGCCCAACAAUAA ..(((((((((((......)))......(((..(((((((((((.....)))).)))))))...)))--------))))))))((((((((((...)))..)))))))..... ( -48.20) >DroSec_CAF1 14170 112 + 1 CUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGCUUAUGUGGCUCUACUUGCAGGUAGU-AGGCGCGUUCCGCAAGCCCAACAAUAA .....((((.(((.(((.(((((.(((((((..(((((((((((.....)))).))))))).))).)))).))))))))))).))-))(((.....))).............. ( -48.00) >DroSim_CAF1 6780 112 + 1 CUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGNNNNNNNNNNNNNNNNNNNNNNNNNN-NNNNNNNNNNNNNNNNNNNNNNNNNNN .(((((((((....)).))))))).(((.....)))((((((((.....))))))))............................-........................... ( -22.20) >consensus CUUAUCUGCAGCCGUGCCAGGUAAAGCCCAUUUGGCGGGAUCCGUUCUGCGGGUUCCCGCUU_UGUG________UGCAGGUAGU__GGCGCGUUCCGCAAGCCCAACAAUAA .(((((((((....)).))))))).(((.....)))((((((((.....))))))))..........................(((((((...........)))))))..... (-24.95 = -26.20 + 1.25)

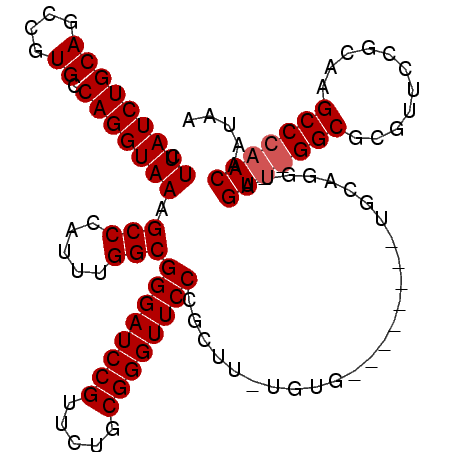
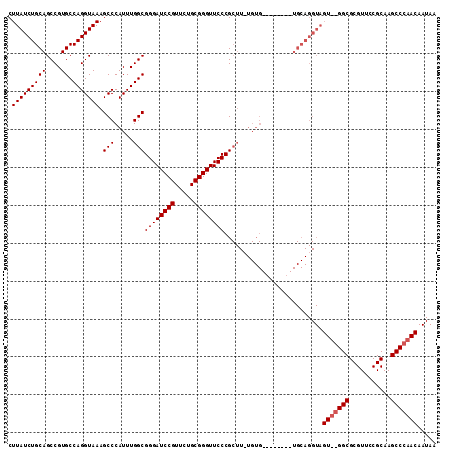
| Location | 13,481,388 – 13,481,493 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 65.09 |
| Mean single sequence MFE | -32.20 |
| Consensus MFE | -20.73 |
| Energy contribution | -23.03 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.64 |
| SVM decision value | 2.24 |
| SVM RNA-class probability | 0.990982 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13481388 105 - 22224390 UUAUUGUUGGGCUUGCGGAACGCGCCCAACUACCUGCA--------CACAAAAGCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAG .....(((((((..(((...))))))))))...(((((--------..(......((((..(((.....))).))))((((...((......))))))..)..)))))..... ( -38.00) >DroSec_CAF1 14170 112 - 1 UUAUUGUUGGGCUUGCGGAACGCGCCU-ACUACCUGCAAGUAGAGCCACAUAAGCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAG ...(((((((((..(((...)))))))-)...(((((..((((((((.(((..((((((..(((.....))).))))))...)))))))))))...))).))..))))..... ( -42.00) >DroSim_CAF1 6780 112 - 1 NNNNNNNNNNNNNNNNNNNNNNNNNNN-NNNNNNNNNNNNNNNNNNNNNNNNNNCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAG ...........................-...........................((((..(((.....))).))))((((...((......))))))............... ( -16.60) >consensus UUAUUGUUGGGCUUGCGGAACGCGCC__ACUACCUGCA________CACA_AAGCGGGAACCCGCAGAACGGAUCCCGCCAAAUGGGCUUUACCUGGCACGGCUGCAGAUAAG .....(((((((..((.....)))))))))...((((..................((((..(((.....))).))))((((...((......))))))......))))..... (-20.73 = -23.03 + 2.31)


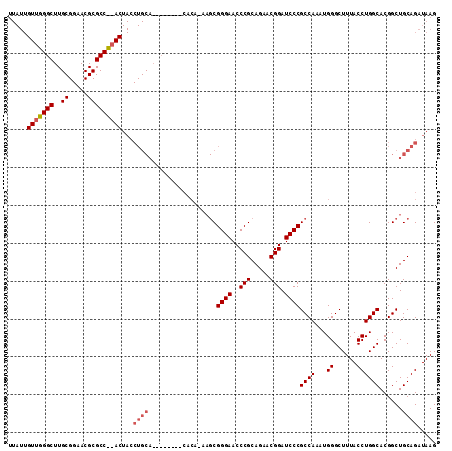
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:41:27 2006