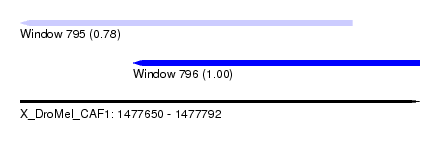
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,477,650 – 1,477,792 |
| Length | 142 |
| Max. P | 0.999758 |

| Location | 1,477,650 – 1,477,768 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.71 |
| Mean single sequence MFE | -28.03 |
| Consensus MFE | -22.88 |
| Energy contribution | -22.44 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.775170 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
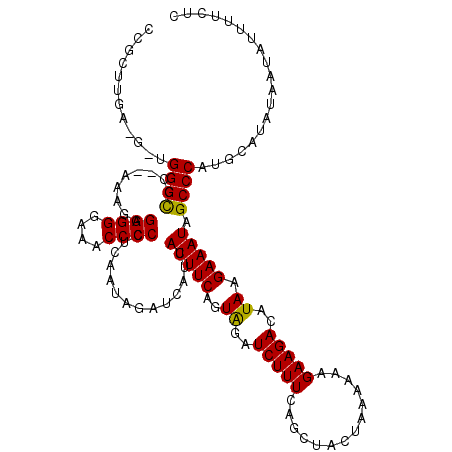
>X_DroMel_CAF1 1477650 118 - 22224390 CCGCUUGAAGGUGGGCGC--AAAGAGGGGGAAACCCCCUCAAUAGAUCAUCAUUUCAGUAGAUCUUUCAGCUACUAAAAAAGAAGACAUAAGAAAUAGCCCAUGCAUAUAAUAUUUUCUC ..........((((((..--...(((((((...)))))))..((..((.((.(((.(((((.........)))))..))).)).))..)).......))))))................. ( -29.10) >DroSec_CAF1 13066 118 - 1 CCGCUUGAUGUUGGGUGC--AAAGAGGGGGAAACCCCCUCAAUAGAUCAUCAUUUCAGUAGAUCUUUCAGCUACUAAAAAAGAAGACAUAAGAAAUAGCCCAUGCAUAUAAUAUUUUCUC .......(((((((((..--...(((((((...)))))))..((..((.((.(((.(((((.........)))))..))).)).))..)).......))))).))))............. ( -27.90) >DroEre_CAF1 14241 115 - 1 CUCCUUAA----GGGCACAGAAAAGGGGGGAAACCCCAGCAAUAGAUCAUAAUUUCAGUGGAUCUUUCAGCUACCAA-AAAGAAGAAAUAAGAAAUAGCCCAUGCAUACAAUAUUUUCUC ........----((((.........((((....))))(((...(((((...((....)).)))))....))).....-...................))))................... ( -27.10) >consensus CCGCUUGA_G_UGGGCGC__AAAGAGGGGGAAACCCCCUCAAUAGAUCAUCAUUUCAGUAGAUCUUUCAGCUACUAAAAAAGAAGACAUAAGAAAUAGCCCAUGCAUAUAAUAUUUUCUC ............((((.........((((....))))..............(((((..((..(((((..............)))))..)).))))).))))................... (-22.88 = -22.44 + -0.44)

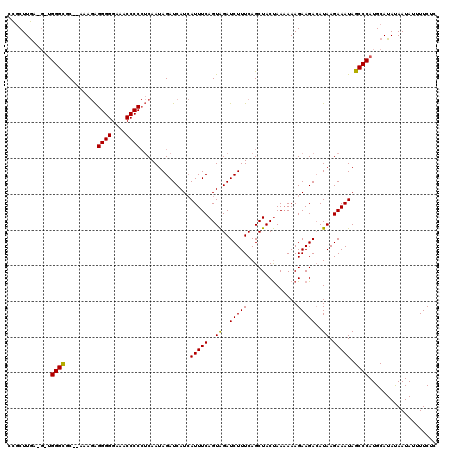
| Location | 1,477,690 – 1,477,792 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.14 |
| Mean single sequence MFE | -27.50 |
| Consensus MFE | -19.24 |
| Energy contribution | -18.80 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.70 |
| SVM decision value | 4.02 |
| SVM RNA-class probability | 0.999758 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
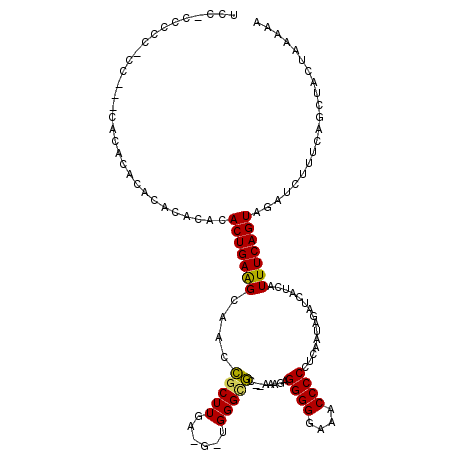
>X_DroMel_CAF1 1477690 102 - 22224390 ----------------GCACACACACACACACUGAAGCAACCGCUUGAAGGUGGGCGC--AAAGAGGGGGAAACCCCCUCAAUAGAUCAUCAUUUCAGUAGAUCUUUCAGCUACUAAAAA ----------------((............((((((((..(((((....)))))..))--...(((((((...))))))).............)))))).((....)).))......... ( -31.20) >DroSec_CAF1 13106 118 - 1 UCCCCCCCCUCCCCACACACACACACACACACUGAGGAAACCGCUUGAUGUUGGGUGC--AAAGAGGGGGAAACCCCCUCAAUAGAUCAUCAUUUCAGUAGAUCUUUCAGCUACUAAAAA ....(((((((....(((...((.(((((...((.(....)))..)).))))).))).--...)))))))......................(((.(((((.........))))).))). ( -24.90) >DroEre_CAF1 14281 115 - 1 UCCACCCCCACCAUCCACACUCACACACAAACUGAAGCAGCUCCUUAA----GGGCACAGAAAAGGGGGGAAACCCCAGCAAUAGAUCAUAAUUUCAGUGGAUCUUUCAGCUACCAA-AA ............((((((..................((.((((.....----)))).........((((....)))).))....((........)).))))))..............-.. ( -26.40) >consensus UCC_CCCCC_CC___CACACACACACACACACUGAAGCAACCGCUUGA_G_UGGGCGC__AAAGAGGGGGAAACCCCCUCAAUAGAUCAUCAUUUCAGUAGAUCUUUCAGCUACUAAAAA ..............................(((((((....(((((......)))))........((((....))))...............)))))))..................... (-19.24 = -18.80 + -0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:26 2006