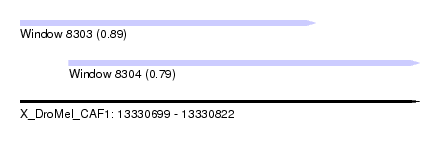

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,330,699 – 13,330,822 |
| Length | 123 |
| Max. P | 0.886637 |

| Location | 13,330,699 – 13,330,790 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 82.65 |
| Mean single sequence MFE | -28.12 |
| Consensus MFE | -23.64 |
| Energy contribution | -23.50 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886637 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13330699 91 + 22224390 UCACGUGUGA------------------GUGGAGUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA-GCGAGCUACUGCAAAUAAUUGCAAAGU ....((.((.------------------.((..((.((((((..(((..((((....))))..)))..)))))).))..)).-.)).))...(((((....))))).... ( -28.40) >DroPse_CAF1 93933 76 + 1 ---------------------------------GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAGGCACUUGGCGACGA-GCAAGCUACUGCAAAUAAUUGCAAAGU ---------------------------------((.((((((..(((..((((....))))..)))..)))))).)).....-....(((..(((((....))))).))) ( -24.10) >DroSim_CAF1 35080 91 + 1 UCACGUGUGA------------------GUGGAGUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA-GCGAGCUACUGCAAAUAAUUGCAAAGU ....((.((.------------------.((..((.((((((..(((..((((....))))..)))..)))))).))..)).-.)).))...(((((....))))).... ( -28.40) >DroEre_CAF1 34666 91 + 1 UCACGUGUGA------------------GUGGAGUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA-GCGAGCUACUGCAAAUAAUUGCAAAGU ....((.((.------------------.((..((.((((((..(((..((((....))))..)))..)))))).))..)).-.)).))...(((((....))))).... ( -28.40) >DroAna_CAF1 19437 110 + 1 CCCCUUGUGACUAAGCCACUGGAUACUGGUUGAGUGAAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGGGGCGAGCUACUGCAAAUAAUUGCAAAGU ((((((((.(((.(((((........))))).))).((((((..(((..((((....))))..)))..)))))).)))).))))...(((..(((((....))))).))) ( -34.80) >DroPer_CAF1 94330 78 + 1 -------------------------------GUGUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAGGCACUUGGCGACGA-GCAAGCUACUGCAAAUAAUUGCAAAGU -------------------------------((((.((((((..(((..((((....))))..)))..)))))).)).))..-....(((..(((((....))))).))) ( -24.60) >consensus UCACGUGUGA__________________GUGGAGUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA_GCGAGCUACUGCAAAUAAUUGCAAAGU .................................((.((((((..(((..((((....))))..)))..)))))).))..........(((..(((((....))))).))) (-23.64 = -23.50 + -0.14)



| Location | 13,330,714 – 13,330,822 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 90.31 |
| Mean single sequence MFE | -34.52 |
| Consensus MFE | -29.78 |
| Energy contribution | -29.28 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792282 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13330714 108 + 22224390 GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA-GCGAGCUACUGCAAAUAAUUGCAAAGUCUU-UGGCCAAGUCUUGG-CAGCCACUGCUGCUG ((.((((((..(((..((((....))))..)))..)))))).))(((..-((.((..(((((((....))))..)))..)-).))...)))..((-((((....)))))). ( -37.70) >DroPse_CAF1 93933 97 + 1 GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAGGCACUUGGCGACGA-GCAAGCUACUGCAAAUAAUUGCAAAGUCUUUUGGCCAAGUCUUGG-------------CUG ((.((((((..(((..((((....))))..)))..)))))).))(((.(-(....))..(((((....)))))..)))....(((((.....)))-------------)). ( -32.70) >DroEre_CAF1 34681 105 + 1 GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA-GCGAGCUACUGCAAAUAAUUGCAAAGUCUU-UGGCCAAGUCUUGG-CAGCCGCU---GCUG ((.((((((..(((..((((....))))..)))..)))))).))....(-(((.(((..(((((....))))).......-..((((.....)))-))))))))---.... ( -37.40) >DroYak_CAF1 34693 105 + 1 GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA-GCGAGCUACUGCAAAUAAUUGCAAAGUCUU-UGGCCAAGUCUUGA-CAGCCACU---GCUG ((.((((((..(((..((((....))))..)))..)))))).))(((..-((.((..(((((((....))))..)))..)-).))...)))....-((((....---)))) ( -32.30) >DroAna_CAF1 19470 107 + 1 GUGAAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGGGGCGAGCUACUGCAAAUAAUUGCAAAGUCUC-UGGCAAAGUCUUGGGCUUCCACU---GCUG ((.((((((..(((..((((....))))..)))..)))))).))(((...((.((..(((((((....))))..)))..)-).))...)))...(((.......---))). ( -34.30) >DroPer_CAF1 94332 97 + 1 GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAGGCACUUGGCGACGA-GCAAGCUACUGCAAAUAAUUGCAAAGUCUUUUGGCCAAGUCUUGG-------------CUG ((.((((((..(((..((((....))))..)))..)))))).))(((.(-(....))..(((((....)))))..)))....(((((.....)))-------------)). ( -32.70) >consensus GUGGAGUGCAGGCCUCGUAAAAGUUUAUAUGGCAAGCACUUGGCGACGA_GCGAGCUACUGCAAAUAAUUGCAAAGUCUU_UGGCCAAGUCUUGG_CAGCCACU___GCUG ((.((((((..(((..((((....))))..)))..)))))).))(((...(((((((..(((((....))))).)).)))...))...))).................... (-29.78 = -29.28 + -0.50)

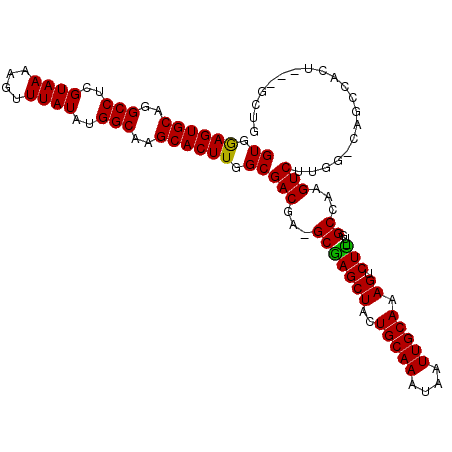

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:16 2006