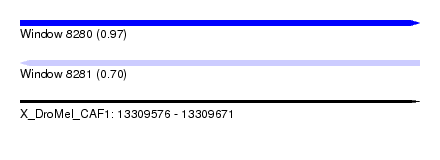

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,309,576 – 13,309,671 |
| Length | 95 |
| Max. P | 0.970032 |

| Location | 13,309,576 – 13,309,671 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 91.56 |
| Mean single sequence MFE | -9.64 |
| Consensus MFE | -8.61 |
| Energy contribution | -8.61 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.970032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
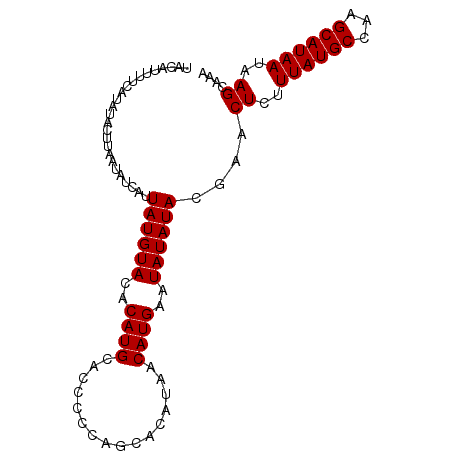
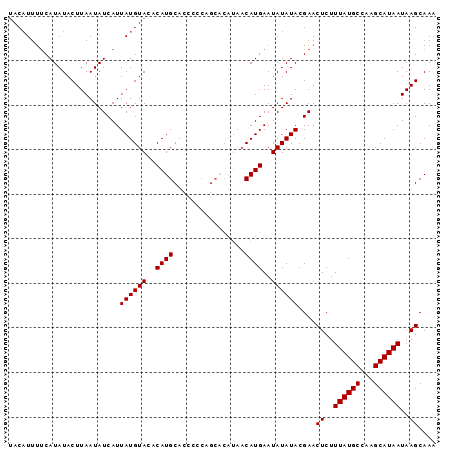
>X_DroMel_CAF1 13309576 95 + 22224390 UAC-UUUUCAUAUACUUAAUAUCAUUAUGUACACAUGCACCCCCAGCACAUAACAUGAAUAUAUACGAACUCUUUAUGCCAAGCAUAAUAAGCAAA ...-..............((((..((((((.....(((.......)))....))))))..)))).....((..((((((...))))))..)).... ( -10.30) >DroSec_CAF1 22369 96 + 1 UACAUUUUCAUAUACUUAAUAUCAUUAUGUACACAUGCACCCCCCGCCCAUAACAUGAAUAUAUACGAACUCUUUAUGCCAAGCAUAAUAAGCAAA ..................((((..((((((......((.......)).....))))))..)))).....((..((((((...))))))..)).... ( -8.70) >DroSim_CAF1 14569 96 + 1 UACAUUUUCAUAUACUUAAUAUCAUUAUGUACACAUGCACCCCCAGCACAUAACAUGAAUAUAUACGAACUCUUUAUGCCAAGCAUAAUAAGCAAA ..................((((..((((((.....(((.......)))....))))))..)))).....((..((((((...))))))..)).... ( -10.30) >DroEre_CAF1 14138 96 + 1 UACAUUCCCAUAUACUUAAUAUCAUUAUGUACACAUGCACUCCCAGCACAUAACAUGAAUAUAUACGAACUCUUUAUGCCAAGCAUAAUAAGCAAA ..................((((..((((((.....(((.......)))....))))))..)))).....((..((((((...))))))..)).... ( -10.30) >DroYak_CAF1 13321 82 + 1 CACAUUUUCAUAUACUUAAUAUCAUUAUGUACACAUGCA--------------CAUGAAUAUAUACGAACUCUUUAUGCCAAGCAUAAUAAGCAAA ..................((((..((((((.(....).)--------------)))))..)))).....((..((((((...))))))..)).... ( -8.60) >consensus UACAUUUUCAUAUACUUAAUAUCAUUAUGUACACAUGCACCCCCAGCACAUAACAUGAAUAUAUACGAACUCUUUAUGCCAAGCAUAAUAAGCAAA .........................((((((..((((................))))..))))))....((..((((((...))))))..)).... ( -8.61 = -8.61 + 0.00)

| Location | 13,309,576 – 13,309,671 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 91.56 |
| Mean single sequence MFE | -19.69 |
| Consensus MFE | -16.14 |
| Energy contribution | -16.48 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.703436 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
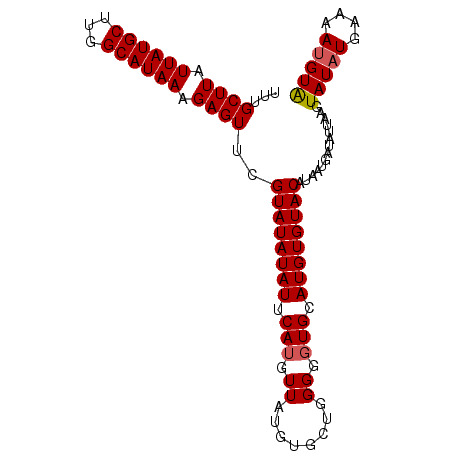
>X_DroMel_CAF1 13309576 95 - 22224390 UUUGCUUAUUAUGCUUGGCAUAAAGAGUUCGUAUAUAUUCAUGUUAUGUGCUGGGGGUGCAUGUGUACAUAAUGAUAUUAAGUAUAUGAAAA-GUA ...((((.((((((...)))))).))))((((((((......(((((((((.............)))))))))........))))))))...-... ( -20.36) >DroSec_CAF1 22369 96 - 1 UUUGCUUAUUAUGCUUGGCAUAAAGAGUUCGUAUAUAUUCAUGUUAUGGGCGGGGGGUGCAUGUGUACAUAAUGAUAUUAAGUAUAUGAAAAUGUA ...((((.((((((...)))))).))))..((((((((.(((.((........)).))).))))))))..............(((((....))))) ( -18.60) >DroSim_CAF1 14569 96 - 1 UUUGCUUAUUAUGCUUGGCAUAAAGAGUUCGUAUAUAUUCAUGUUAUGUGCUGGGGGUGCAUGUGUACAUAAUGAUAUUAAGUAUAUGAAAAUGUA ...((((.((((((...)))))).))))((((((((......(((((((((.............)))))))))........))))))))....... ( -20.36) >DroEre_CAF1 14138 96 - 1 UUUGCUUAUUAUGCUUGGCAUAAAGAGUUCGUAUAUAUUCAUGUUAUGUGCUGGGAGUGCAUGUGUACAUAAUGAUAUUAAGUAUAUGGGAAUGUA ...((((.((((((...)))))).))))((.((((((((...(((((((((.............))))))))).......)))))))).))..... ( -20.62) >DroYak_CAF1 13321 82 - 1 UUUGCUUAUUAUGCUUGGCAUAAAGAGUUCGUAUAUAUUCAUG--------------UGCAUGUGUACAUAAUGAUAUUAAGUAUAUGAAAAUGUG ...((((.((((((...)))))).))))...(((((.((((((--------------(((..((((........))))...))))))))).))))) ( -18.50) >consensus UUUGCUUAUUAUGCUUGGCAUAAAGAGUUCGUAUAUAUUCAUGUUAUGUGCUGGGGGUGCAUGUGUACAUAAUGAUAUUAAGUAUAUGAAAAUGUA ...((((.((((((...)))))).))))..((((((((.(((.((........)).))).))))))))..............(((((....))))) (-16.14 = -16.48 + 0.34)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:55 2006