
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,307,426 – 13,307,656 |
| Length | 230 |
| Max. P | 0.982093 |

| Location | 13,307,426 – 13,307,524 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 82.20 |
| Mean single sequence MFE | -21.88 |
| Consensus MFE | -19.21 |
| Energy contribution | -18.53 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938584 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13307426 98 - 22224390 AUACUGUCCAGUAAUGGCCAUAAUGGAAACGGUUAUAAACGAAAUAAUAGCCGAAACAAUAGUU---------------GAAAAUAUAGUGGCUGUGAAGUCGUAAUUAUUUA ..((((((((....)))((.....))..))))).....((((....(((((((.(((....)))---------------..........)))))))....))))......... ( -19.50) >DroSec_CAF1 17086 113 - 1 AUACUGUCCAGUAACGGCCAUAAUGGAAACGGUUAUAAACGAAAUAAUAGCCGAAACAAUAGUUGAGGGAACAAAACUUGAAAAUAUAGUGGCUGUGAAUUCAAAAUUAUUUA .((((....))))((((((((.(((....(((((((..........)))))))...((..((((..(....)..))))))....))).))))))))................. ( -23.80) >DroSim_CAF1 11063 113 - 1 AUACUGUCCAGUAAUAGCCAUAAUGGAAACGGUUAUAAACGAAAUAAUAGCCGAAACAAUAGUUGAGGGAACGAAACUUGAAAAUAUAGUGGCUGUGAAUUCAAAACUAUUUA .((((....))))((((((((.(((....(((((((..........)))))))...((..((((..(....)..))))))....))).))))))))................. ( -23.60) >DroYak_CAF1 11440 102 - 1 AUACUGUCCAGUAAUGGCCAUAAUGGAAACGCUUAUAAGCGACAUAAUAUGCGAAACAAUAGUUGAGAU----AUAGUUGAG-AUAUAGUGGCUAUUAAG------UUAUUUA ...........((((((((((.(((....((((....)))).((...(((((..(((....)))..).)----)))..))..-.))).))))))))))..------....... ( -20.60) >consensus AUACUGUCCAGUAAUGGCCAUAAUGGAAACGGUUAUAAACGAAAUAAUAGCCGAAACAAUAGUUGAGGG____AAACUUGAAAAUAUAGUGGCUGUGAAGUCAAAAUUAUUUA .((((....))))((((((((.(((....(((((((..........))))))).(((....)))....................))).))))))))................. (-19.21 = -18.53 + -0.69)



| Location | 13,307,524 – 13,307,626 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 98.04 |
| Mean single sequence MFE | -28.82 |
| Consensus MFE | -27.18 |
| Energy contribution | -27.42 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.982093 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13307524 102 + 22224390 CCCUUGAGCUUCUGCUGUGUCACCGAGUGCAUUUUUCGCUUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA ......(((....))).....(((...(((.((((((((((((.(((((......))))).(((........))).)).)))))))))).)))...)))... ( -29.40) >DroSec_CAF1 17199 102 + 1 CCCUUGAGCUUCUGCUGUGUCACCGAGUGCAUUUUUCGCUUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA ......(((....))).....(((...(((.((((((((((((.(((((......))))).(((........))).)).)))))))))).)))...)))... ( -29.40) >DroSim_CAF1 11176 102 + 1 CCCUUGAGCUUCUGCUGUGUCACCGAGUGCAUUUUUCGCUUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA ......(((....))).....(((...(((.((((((((((((.(((((......))))).(((........))).)).)))))))))).)))...)))... ( -29.40) >DroEre_CAF1 12067 101 + 1 CCCUUGAGCUUCUGCUGCGUCACCGAGUGCAUUUUUCGCUUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAA-AAGCGAGAGGUAAA .((((..((...(((..((....))...)))((((((((((((.(((((......))))).(((........))).)).))))))))-))))..)))).... ( -29.10) >DroYak_CAF1 11542 102 + 1 CCCUUGAUCUGCUGCUGUGUCAUCGAGUGCAUUUUUCGCUUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA ((((((((..((......)).))))))(((.((((((((((((.(((((......))))).(((........))).)).)))))))))).)))...)).... ( -26.80) >consensus CCCUUGAGCUUCUGCUGUGUCACCGAGUGCAUUUUUCGCUUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA ......(((....))).....(((...(((.((((((((((((.(((((......))))).(((........))).)).)))))))))).)))...)))... (-27.18 = -27.42 + 0.24)

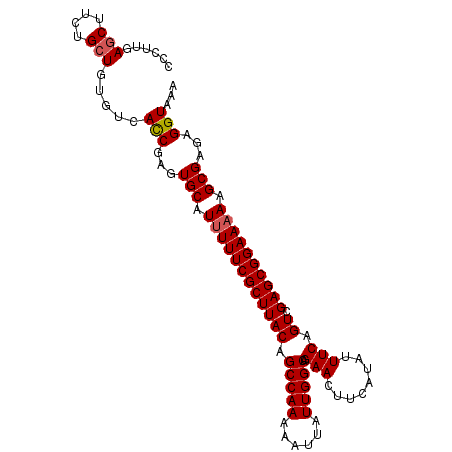
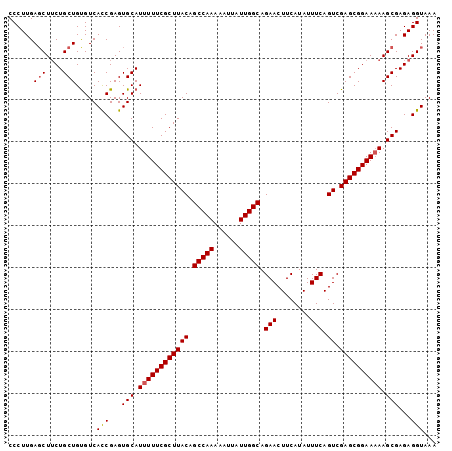
| Location | 13,307,563 – 13,307,656 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 97.40 |
| Mean single sequence MFE | -20.42 |
| Consensus MFE | -19.24 |
| Energy contribution | -19.68 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830518 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13307563 93 + 22224390 UUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAAGAACUUUUGCAAAUGUCAUGCAGGUGAUUG .....(((((......)))))..............((((((.(((((....(((((((........))))))).....)).)))...)))))) ( -22.10) >DroSec_CAF1 17238 92 + 1 UUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA-AAAUUUUGCAAAUGUCAUGCAGGUGAUUG .....(((((......)))))..............((((((.(((((....((((((......-...)))))).....)).)))...)))))) ( -17.90) >DroSim_CAF1 11215 92 + 1 UUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA-AACUUUUGCAAAUGUCAUGCAGGUGAUUG .....(((((......)))))..............((((((.(((((....(((((((.....-..))))))).....)).)))...)))))) ( -21.80) >DroEre_CAF1 12106 91 + 1 UUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAA-AAGCGAGAGGUAAA-AACGAUUGCAAAUGUCAUGCAGGUGGUUG .....(((((......)))))...(((((.....((....))((....-..))..)))))...-..((((..(...((.....)).)..)))) ( -18.50) >DroYak_CAF1 11581 92 + 1 UUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA-AACUUUUGCAAAUGUCAUGCAGGUGAUUG .....(((((......)))))..............((((((.(((((....(((((((.....-..))))))).....)).)))...)))))) ( -21.80) >consensus UUACAGCCAAAAAUUAUUGGCAGAACUUCAUAUUUCAGUCGAGCGGAAAAAGCGAGAGGUAAA_AACUUUUGCAAAUGUCAUGCAGGUGAUUG .....(((((......)))))..............((((((.(((((....(((((((........))))))).....)).)))...)))))) (-19.24 = -19.68 + 0.44)


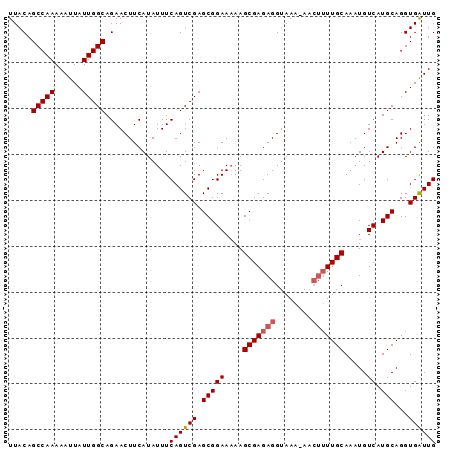
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:53 2006