
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,296,599 – 13,296,735 |
| Length | 136 |
| Max. P | 0.911179 |

| Location | 13,296,599 – 13,296,703 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 97.60 |
| Mean single sequence MFE | -23.50 |
| Consensus MFE | -23.31 |
| Energy contribution | -23.38 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911179 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13296599 104 + 22224390 UUUUAUUGUACGACAAAAUAUUUCUUUAUUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCC ...........((((((................((((.((.(((((....))))))).))))((((((((((((....)).))))))...)))).))))))... ( -24.40) >DroSec_CAF1 11846 104 + 1 UUUUAUUGUACGACAAAAUAUUUCUUUAUUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCC ...........((((((................((((.((.(((((....))))))).))))((((((((((((....)).))))))...)))).))))))... ( -24.40) >DroSim_CAF1 5753 104 + 1 UUUUAUUGUACGACAAAAUAUUUCUUUAUUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCC ...........((((((................((((.((.(((((....))))))).))))((((((((((((....)).))))))...)))).))))))... ( -24.40) >DroEre_CAF1 6739 104 + 1 UUUUAUUGUACUACAAAAUAUUUCUUUAUUAUCGUGAGUGAGUUGAAUUUUCAGCCGAUCACGUGAACAACUCAGCCUUGAAGUUGUGUUUCGCAUUUGUUUCC ............(((((................((((.((.(((((....))))))).))))(((((..((.((((......)))).))))))).))))).... ( -20.80) >consensus UUUUAUUGUACGACAAAAUAUUUCUUUAUUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCC ...........((((((................((((.((.(((((....))))))).))))((((((((((((....)).))))))...)))).))))))... (-23.31 = -23.38 + 0.06)



| Location | 13,296,627 – 13,296,735 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 94.76 |
| Mean single sequence MFE | -20.65 |
| Consensus MFE | -20.69 |
| Energy contribution | -20.50 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.53 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13296627 108 + 22224390 UUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCCGUUCUUAUUUUUAUAUUUAUAUAUGUACCUAU .....((((.((.(((((....))))))).))))((((((((((((....)).))))))...)))).....................(((((......)))))..... ( -21.50) >DroSec_CAF1 11874 102 + 1 UUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCCGUUCUUAUUUUUAU--U----UAUAUACCUAU .....((((.((.(((((....))))))).))))((((((((((((....)).))))))...))))........................--.----........... ( -21.10) >DroSim_CAF1 5781 102 + 1 UUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCCGUUCUUAUUUUUAU--U----UAUAUACCUAU .....((((.((.(((((....))))))).))))((((((((((((....)).))))))...))))........................--.----........... ( -21.10) >DroEre_CAF1 6767 102 + 1 UUAUCGUGAGUGAGUUGAAUUUUCAGCCGAUCACGUGAACAACUCAGCCUUGAAGUUGUGUUUCGCAUUUGUUUCCGUUCUUAUUUUUAU--U----UAUAUACCUAU .....((((.((.(((((....))))))).))))(((((..((.((((......)))).)))))))........................--.----........... ( -18.90) >consensus UUAUCGUGAGUGAGUUGAACUUUCAGCCGAUCACGCGAACAACUCAACCUUGAAGUUGUAUUUCGCAUUUGUUUCCGUUCUUAUUUUUAU__U____UAUAUACCUAU .....((((.((.(((((....))))))).))))((((((((((((....)).))))))...)))).......................................... (-20.69 = -20.50 + -0.19)

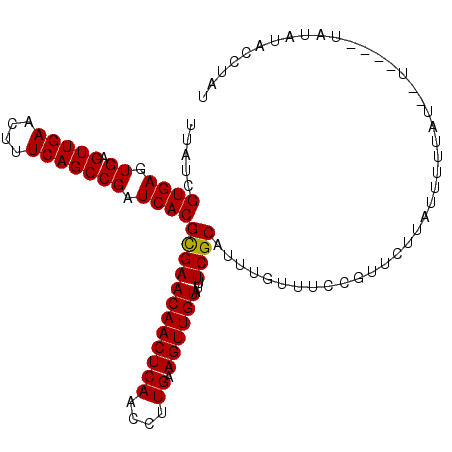

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:50 2006