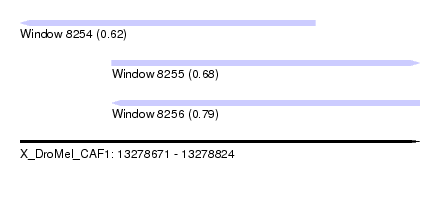
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,278,671 – 13,278,824 |
| Length | 153 |
| Max. P | 0.791151 |

| Location | 13,278,671 – 13,278,784 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.26 |
| Mean single sequence MFE | -40.14 |
| Consensus MFE | -30.18 |
| Energy contribution | -30.74 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.624243 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13278671 113 - 22224390 CAUUG--UUCGAGUGCGAGUGAGAUGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGCACUUC-----CGACGUCGUCCUGGGCCAAGGAAGUUGUAG ..(..--((((....))))..)..((((((((((((((((...((((((((..........))))))))))))))))..))))))-----))(((...((((......))))...))).. ( -38.80) >DroSec_CAF1 24882 113 - 1 CUUCG--CUCGAGUGCGAGUGAGACGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGCACUUC-----CGACGUCGUCCUGGGCCAAGGAAGUUGUAG .((((--((((....)))))))).((((((((((((((((...((((((((..........))))))))))))))))..))))))-----))(((...((((......))))...))).. ( -46.50) >DroSim_CAF1 23501 113 - 1 CUUUG--CUCGAGUGCGAGUGAGACGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGCACUUC-----CGACGUCGUCCUGGGCCAAGGAAGUUGUAG .((..--((((....))))..)).((((((((((((((((...((((((((..........))))))))))))))))..))))))-----))(((...((((......))))...))).. ( -44.10) >DroEre_CAF1 24698 112 - 1 CUGUACACGCGAGUGCGAGUGAGGCG--AGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUU-UUUACUCAUUUCCGCUGGCACUUC-----CGACGUCGUCCUGGGCCAAGGAAGUUGUAG ((.(((.(((....))).)))))(((--(((.((((((((...((((((((.........-)))))))))))))))).)).((((-----(...(((......)))...))))))))).. ( -37.70) >DroYak_CAF1 24365 108 - 1 ---------CGGGUGCGAGUGAGACG--GAUGAGUGGGACUUUUGAGUGAGUUGUAGGUU-UUUACUCAUUUCCGCUGGCACUUCCGUUCCGUCGUCGUCCUGGGCCAAGGAAGUUGUAG ---------((((.((((....((((--((..(((((((....((((((((.........-)))))))).)))))))((.....))..)))))).))))))))................. ( -33.60) >consensus CUUUG__CUCGAGUGCGAGUGAGACGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGCACUUC_____CGACGUCGUCCUGGGCCAAGGAAGUUGUAG .............(((((.(..((((((((((((((((((...((((((((..........))))))))))))))))..))))))........)))).((((......))))).))))). (-30.18 = -30.74 + 0.56)



| Location | 13,278,706 – 13,278,824 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.88 |
| Mean single sequence MFE | -26.75 |
| Consensus MFE | -17.56 |
| Energy contribution | -18.79 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.679759 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13278706 118 + 22224390 GCCAGCGGAAAUGAGUAAAAAACCUACAACUCACUCAAAAUUCCCGCUCACUCCCAUCUCACUCGCACUCGAA--CAAUGGGAAAUUCCGUUUCGAGUGAAAUUUUGUGCCACAAACGAA ...(((((.(((((((................)))))...)).)))))..............(((((((((((--..(((((....)))))))))))))....((((.....))))))). ( -24.99) >DroSec_CAF1 24917 117 + 1 GCCAGCGGAAAUGAGUAAAAAACCUACAACUCACUCAAAAUUCCCGCUCACUCCCGUCUCACUCGCACUCGAG--CGAAGGGAAAUUCCGUUUCGAGUGAAAUU-CGGGCCACACACGAA ...(((((.(((((((................)))))...)).)))))....((((..((((((((......)--)((((((.....)).))))))))))....-))))........... ( -29.39) >DroSim_CAF1 23536 117 + 1 GCCAGCGGAAAUGAGUAAAAAACCUACAACUCACUCAAAAUUCCCGCUCACUCCCGUCUCACUCGCACUCGAG--CAAAGGGAAAUUCCGUUUCGAGUGGAAUU-CGGGCCACACACGAA ((((((((((.(((((............))))).......(((((((((.....((.......)).....)))--)...))))).))))))).(((((...)))-))))).......... ( -28.20) >DroEre_CAF1 24733 116 + 1 GCCAGCGGAAAUGAGUAAA-AACCUACAACUCACUCAAAAUUCCCGCUCACU--CGCCUCACUCGCACUCGCGUGUACAGGGUAAUUCCACUUCGAGUGAAACU-CAGGCCGCACACGAA (((.((((.(((((((...-............)))))...)).))))(((((--((....((.(((....))).))...(((....)))....)))))))....-..))).......... ( -29.86) >DroYak_CAF1 24405 97 + 1 GCCAGCGGAAAUGAGUAAA-AACCUACAACUCACUCAAAAGUCCCACUCAUC--CGUCUCACUCGCACCCG-------------------UUCCGAGUGAAACU-CAGGCCACACACGAA (((.(((((..(((((...-.........................)))))))--))).(((((((......-------------------...)))))))....-..))).......... ( -21.29) >consensus GCCAGCGGAAAUGAGUAAAAAACCUACAACUCACUCAAAAUUCCCGCUCACUCCCGUCUCACUCGCACUCGAG__CAAAGGGAAAUUCCGUUUCGAGUGAAAUU_CGGGCCACACACGAA ((((((((...(((((................)))))......))))).................((((((((......(((....)))..))))))))........))).......... (-17.56 = -18.79 + 1.23)


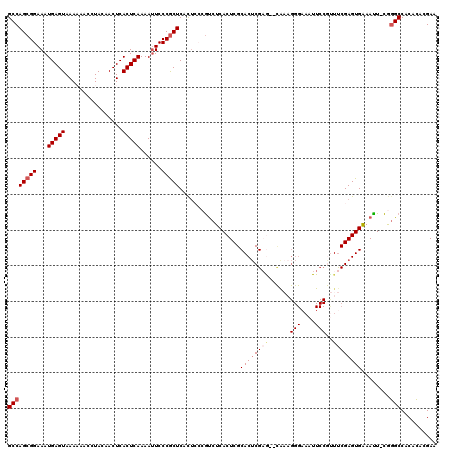
| Location | 13,278,706 – 13,278,824 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.88 |
| Mean single sequence MFE | -39.28 |
| Consensus MFE | -28.49 |
| Energy contribution | -29.01 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.791151 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13278706 118 - 22224390 UUCGUUUGUGGCACAAAAUUUCACUCGAAACGGAAUUUCCCAUUG--UUCGAGUGCGAGUGAGAUGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGC (((((((((((.........))))...)))))))...(((((((.--((((....))))...)))))))((.((((((((...((((((((..........)))))))))))))))).)) ( -37.60) >DroSec_CAF1 24917 117 - 1 UUCGUGUGUGGCCCG-AAUUUCACUCGAAACGGAAUUUCCCUUCG--CUCGAGUGCGAGUGAGACGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGC ...........((((-..(((((((((((..((.....)).)))(--(......)))))))))))))).((.((((((((...((((((((..........)))))))))))))))).)) ( -41.70) >DroSim_CAF1 23536 117 - 1 UUCGUGUGUGGCCCG-AAUUCCACUCGAAACGGAAUUUCCCUUUG--CUCGAGUGCGAGUGAGACGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGC (((((..((((....-....)))).....)))))...((((((..--((((....))))..))..))))((.((((((((...((((((((..........)))))))))))))))).)) ( -41.00) >DroEre_CAF1 24733 116 - 1 UUCGUGUGCGGCCUG-AGUUUCACUCGAAGUGGAAUUACCCUGUACACGCGAGUGCGAGUGAGGCG--AGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUU-UUUACUCAUUUCCGCUGGC ..((....))(((..-.((((((((((.((.((.....))))...(((....))))))))))))).--....((((((((...((((((((.........-))))))))))))))))))) ( -42.40) >DroYak_CAF1 24405 97 - 1 UUCGUGUGUGGCCUG-AGUUUCACUCGGAA-------------------CGGGUGCGAGUGAGACG--GAUGAGUGGGACUUUUGAGUGAGUUGUAGGUU-UUUACUCAUUUCCGCUGGC ..........(((..-.((((((((((.(.-------------------....).)))))))))).--....(((((((....((((((((.........-)))))))).)))))))))) ( -33.70) >consensus UUCGUGUGUGGCCCG_AAUUUCACUCGAAACGGAAUUUCCCUUUG__CUCGAGUGCGAGUGAGACGGGAGUGAGCGGGAAUUUUGAGUGAGUUGUAGGUUUUUUACUCAUUUCCGCUGGC ..........(((....((((((((((............................)))))))))).......((((((((...((((((((..........))))))))))))))))))) (-28.49 = -29.01 + 0.52)


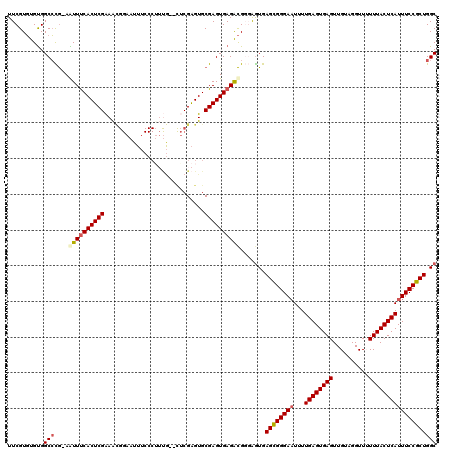
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:33 2006