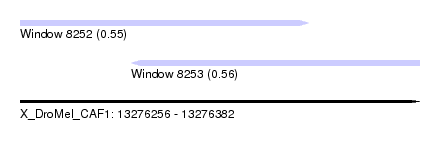
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,276,256 – 13,276,382 |
| Length | 126 |
| Max. P | 0.558110 |

| Location | 13,276,256 – 13,276,347 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 89.47 |
| Mean single sequence MFE | -23.76 |
| Consensus MFE | -16.60 |
| Energy contribution | -17.85 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553178 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13276256 91 + 22224390 UUCGCCGGCUAAAUCGUUGCAAAAUUUUUGUUUGUUUGUUUGCCUAGCUUCGAUUGAAGCCAGUGAGUG-AUUCAAUCGGUUUGUGGACUUU (((((.(((.((((....(((((.......)))))..))))))).....(((((((((..((.....))-.)))))))))...))))).... ( -19.00) >DroSec_CAF1 21688 92 + 1 UUCACUGGCUAAAUCGUGGCAAAAUUUUUGUUUGUUUUUUUGCCAAGCUUCGAUUGAAGCCAGUGAGUGGAUUCAAUCGGUUUGUGGACUUU .(((((((((.(((((((((((((.............)))))))).....)))))..)))))))))(..((((.....))))..)....... ( -28.62) >DroSim_CAF1 21037 92 + 1 UUCGCAGGCUAAAUCGUUGCAAAAUGUUUGUUUGUUUGUUUGCCAAGCUUCGAUUGAAGCCAGUGAGUGGAUUCAAUCGGUUUGUGGACUUU ...((((((.(((.((((....)))))))))))))..(((..(.((((..((((((((.(((.....))).)))))))))))))..)))... ( -25.30) >DroEre_CAF1 22419 86 + 1 UUUGCAGACCAAAUCGUUGAAA-AUUUGUGUUUGUUUGUUUGCCUAGCUUCGAUUGAAGCCCGUG-----AUUCAAUCGGUUUGUGGACUUU ...(((((((((((........-))))).))))))..(((..(..(((..((((((((.......-----.))))))))))).)..)))... ( -22.10) >consensus UUCGCAGGCUAAAUCGUUGCAAAAUUUUUGUUUGUUUGUUUGCCAAGCUUCGAUUGAAGCCAGUGAGUG_AUUCAAUCGGUUUGUGGACUUU ((((((((((...................(((((.........)))))...(((((((.(((.....))).))))))))))))))))).... (-16.60 = -17.85 + 1.25)



| Location | 13,276,291 – 13,276,382 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 93.10 |
| Mean single sequence MFE | -18.07 |
| Consensus MFE | -14.21 |
| Energy contribution | -15.78 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558110 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13276291 91 - 22224390 UGUUAGCAAGUUUUCUAUAAGAAUUUAAGUACUCCAAAGUCCACAAACCGAUUGAAU-CACUCACUGGCUUCAAUCGAAGCUAGGCAAACAA ((((.(((((((((.....)))))))...............................-......((((((((....)))))))))).)))). ( -17.40) >DroSec_CAF1 21723 92 - 1 UGUUAGCAAGUUUUCUAUAAGAAUUUAAGUACUGCAAAGUCCACAAACCGAUUGAAUCCACUCACUGGCUUCAAUCGAAGCUUGGCAAAAAA ......(((((((...............(.(((....))).)......((((((((.(((.....))).)))))))))))))))........ ( -19.40) >DroSim_CAF1 21072 92 - 1 UGUUAGCAAGUUUUCUAUAAGAAUUUAAGUACUUCAAAGUCCACAAACCGAUUGAAUCCACUCACUGGCUUCAAUCGAAGCUUGGCAAACAA ((((.((((((((...............(.(((....))).)......((((((((.(((.....))).)))))))))))))).)).)))). ( -20.80) >DroEre_CAF1 22453 87 - 1 UGUUAGUAAGUUUUCUAUAAGAGUUUAAGUACUACAAAGUCCACAAACCGAUUGAAU-----CACGGGCUUCAAUCGAAGCUAGGCAAACAA ((((.((.(((((.........((((..(.(((....))).)..))))((((((((.-----.......)))))))))))))..)).)))). ( -14.70) >consensus UGUUAGCAAGUUUUCUAUAAGAAUUUAAGUACUACAAAGUCCACAAACCGAUUGAAU_CACUCACUGGCUUCAAUCGAAGCUAGGCAAACAA ((((.((((((((...............(.(((....))).)......((((((((.(((.....))).)))))))))))))).)).)))). (-14.21 = -15.78 + 1.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:30 2006