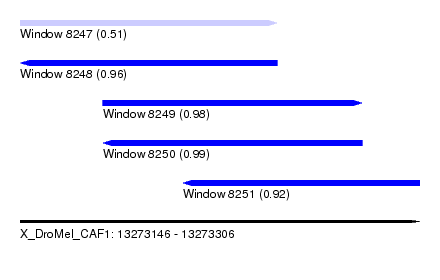

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,273,146 – 13,273,306 |
| Length | 160 |
| Max. P | 0.986002 |

| Location | 13,273,146 – 13,273,249 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 89.35 |
| Mean single sequence MFE | -28.98 |
| Consensus MFE | -22.36 |
| Energy contribution | -22.40 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.77 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.507685 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13273146 103 + 22224390 -------GUGUGUGUUUUGCAUCCUGUUCGUCCUUGCUGCUCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGU -------....((((((.((....(((.((........((((((((((.......)))).....))))))(((((.....)))))....)))))....)).))))))... ( -27.00) >DroSec_CAF1 18592 100 + 1 -------GUGUGU---UGGCAUCCUGCUCGUCCGUGCUGCUCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGU -------((((.(---((((....((.(((........((((((((((.......)))).....))))))(((((.....)))))....)))))....)))))))))... ( -28.30) >DroSim_CAF1 17980 100 + 1 -------GUGUGU---UUGCAUCCUGUUCGUCCUUGCUGCUCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGU -------..((((---(((.((...(((.(((......((((((((((.......)))).....))))))(((((.....))))).....))))))..)))))))))... ( -28.60) >DroEre_CAF1 19345 107 + 1 UGCGCUUGUGUGC---UUGCAUCCUGUUCGUCCUUGCUGCUCGCUCACAUUUCCAGCGACAACAACGAGCACCGCAUUCUGCGGUUUUUCGACAAUUCGUCAGACAUUGU .((((....))))---.........((((((...((.((.(((((.........))))))).))))))))(((((.....))))).....(((.....)))......... ( -27.90) >DroYak_CAF1 18713 107 + 1 GGCACUUGUCUUU---GCACAUCCUGUUCGUCCUUGCUGCUCGUUCACAUUACCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUGCGACAAUUCGUCAGACAUUGU .(((..(((((..---(((......((((((..(((.((((.((.......)).)))).)))..))))))(((((.....)))))..)))(((.....)))))))).))) ( -33.10) >consensus _______GUGUGU___UUGCAUCCUGUUCGUCCUUGCUGCUCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGU .........................((((((..(((.((((.(.........).)))).)))..))))))(((((.....))))).....(((.....)))......... (-22.36 = -22.40 + 0.04)

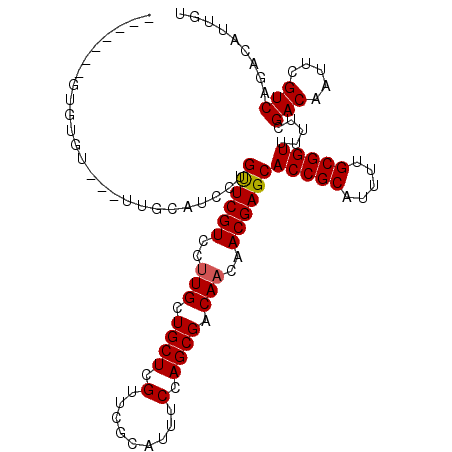

| Location | 13,273,146 – 13,273,249 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 89.35 |
| Mean single sequence MFE | -30.82 |
| Consensus MFE | -27.92 |
| Energy contribution | -27.52 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.960350 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13273146 103 - 22224390 ACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGAGCAGCAAGGACGAACAGGAUGCAAAACACACAC------- .....((((......(((((.....(((((.....)))))((((((.....((((.......))))))))))......)))))...)))).............------- ( -28.00) >DroSec_CAF1 18592 100 - 1 ACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGAGCAGCACGGACGAGCAGGAUGCCA---ACACAC------- .....(((((.(((....)))....(((((.....)))))((((((.....((((.......))))))))))....)))))(.((.....))).---......------- ( -30.10) >DroSim_CAF1 17980 100 - 1 ACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGAGCAGCAAGGACGAACAGGAUGCAA---ACACAC------- ....(((.((.((..((((((....(((((.....)))))((((((.....((((.......))))))))))........)).))))..)))).---)))...------- ( -28.90) >DroEre_CAF1 19345 107 - 1 ACAAUGUCUGACGAAUUGUCGAAAAACCGCAGAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGUGAGCGAGCAGCAAGGACGAACAGGAUGCAA---GCACACAAGCGCA ....((..((.((..((((((....(((((.....)))))..((.(((((((((((.(.....).))))).)))))).)))).))))..)))).---.)).......... ( -31.60) >DroYak_CAF1 18713 107 - 1 ACAAUGUCUGACGAAUUGUCGCAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGUAAUGUGAACGAGCAGCAAGGACGAACAGGAUGUGC---AAAGACAAGUGCC ....((((((((.....)))(((..(((((.....)))))(.(((((.((((.(((.((.......)).))).)))).))))).)......)))---..)))))...... ( -35.50) >consensus ACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGAGCAGCAAGGACGAACAGGAUGCAA___ACACAC_______ .....((((......(((((.....(((((.....)))))((((((.....((((.......))))))))))......)))))...)))).................... (-27.92 = -27.52 + -0.40)

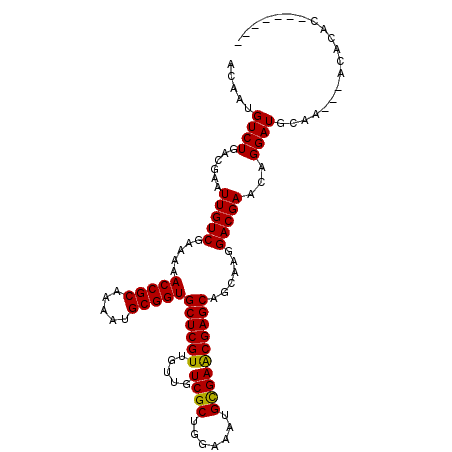
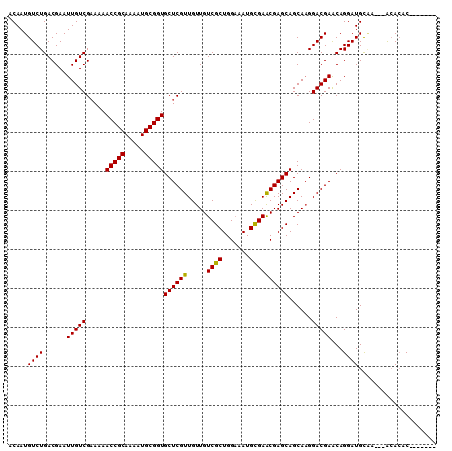
| Location | 13,273,179 – 13,273,283 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 95.58 |
| Mean single sequence MFE | -29.90 |
| Consensus MFE | -26.62 |
| Energy contribution | -26.90 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.983846 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13273179 104 + 22224390 UCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGUUGACACUGUGUCUGAAGUCUGAUGUCUGAUGUCU ..((.(((.......))))).....((((.(((((.....)))))..))))......(((((((((((...(((............))).)))))))))))... ( -30.90) >DroSec_CAF1 18622 104 + 1 UCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGUUGACACCGUGUCUGAAGUCUAAUGUCUGAUGUCU ..((.(((.......))))).....((((.(((((.....)))))..))))......((((((((((((..(((..((....))..)))))))))))))))... ( -31.10) >DroSim_CAF1 18010 104 + 1 UCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGUUGACACUGUGUCUGAAGUCUGAUGUCUGAUGUCU ..((.(((.......))))).....((((.(((((.....)))))..))))......(((((((((((...(((............))).)))))))))))... ( -30.90) >DroEre_CAF1 19382 104 + 1 UCGCUCACAUUUCCAGCGACAACAACGAGCACCGCAUUCUGCGGUUUUUCGACAAUUCGUCAGACAUUGUUGACACUGUGUCUGCAGUCUGAAGUCUGAAGUCU ..(((.........)))(((.....((((.(((((.....)))))..))))........(((((((((((.(((.....))).))))).....)))))).))). ( -28.90) >DroYak_CAF1 18750 104 + 1 UCGUUCACAUUACCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUGCGACAAUUCGUCAGACAUUGUUGACACUGCGUCUGAAGUCUGAUGUCUGAUGUCU (((((.........)))))......((((.(((((.....))))))))).((((...((((((((.((...(((.....))).)).)))))))).....)))). ( -27.70) >consensus UCGUUCGCAUUUCCAGCGACAACAACGAGCACCGCAUUUUGCGGUUUUUCGACAAUUCGUCAGACAUUGUUGACACUGUGUCUGAAGUCUGAUGUCUGAUGUCU (((((.........)))))......((((.(((((.....)))))..))))......(((((((((((...(((............))).)))))))))))... (-26.62 = -26.90 + 0.28)

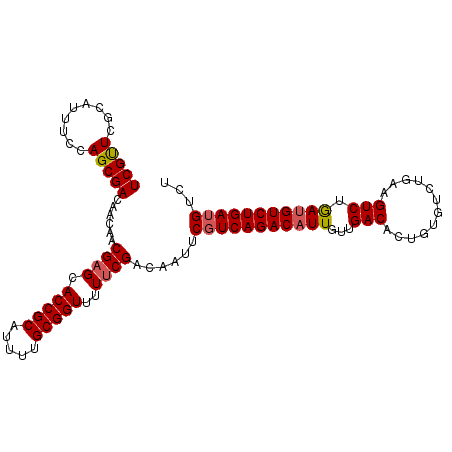

| Location | 13,273,179 – 13,273,283 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 95.58 |
| Mean single sequence MFE | -30.50 |
| Consensus MFE | -27.50 |
| Energy contribution | -27.34 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.986002 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13273179 104 - 22224390 AGACAUCAGACAUCAGACUUCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGA .((((.(.((((.....(.(((((((..((....))..))))))).)...))))((...(((((.....)))))..))).)))).((((.......)))).... ( -30.30) >DroSec_CAF1 18622 104 - 1 AGACAUCAGACAUUAGACUUCAGACACGGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGA .((((.(.((((.....(.(((((((..((....))..))))))).)...))))((...(((((.....)))))..))).)))).((((.......)))).... ( -31.30) >DroSim_CAF1 18010 104 - 1 AGACAUCAGACAUCAGACUUCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGA .((((.(.((((.....(.(((((((..((....))..))))))).)...))))((...(((((.....)))))..))).)))).((((.......)))).... ( -30.30) >DroEre_CAF1 19382 104 - 1 AGACUUCAGACUUCAGACUGCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAGAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGUGAGCGA ...............(..((((....(((((.((((((((..((((.....))))....(((((.....)))))...)))))))).)))))....))))..).. ( -30.60) >DroYak_CAF1 18750 104 - 1 AGACAUCAGACAUCAGACUUCAGACGCAGUGUCAACAAUGUCUGACGAAUUGUCGCAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGUAAUGUGAACGA ..................((((.((.(((((.((((((((...(((.....)))((...(((((.....))))))).)))))))).)))))))....))))... ( -30.00) >consensus AGACAUCAGACAUCAGACUUCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCGGUGCUCGUUGUUGUCGCUGGAAAUGCGAACGA ..........................(((((.((((((((..((((.....))))....(((((.....)))))...)))))))).)))))............. (-27.50 = -27.34 + -0.16)

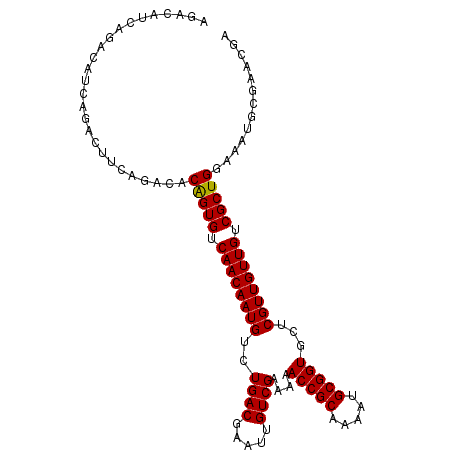
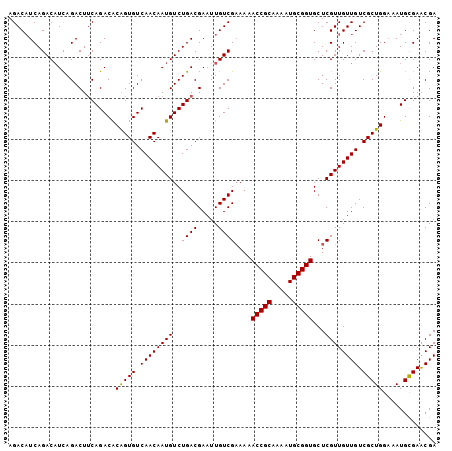
| Location | 13,273,211 – 13,273,306 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 92.83 |
| Mean single sequence MFE | -22.10 |
| Consensus MFE | -18.78 |
| Energy contribution | -18.66 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.918411 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13273211 95 - 22224390 CGACAACG------ACAACGAACCUGCUCAGACAUCAGACAUCAGACUUCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCG ((((((((------....))....((.((((((((..(((((...............)))))....))))))))))..))))))......(((.....))) ( -20.86) >DroSec_CAF1 18654 101 - 1 CGACAACGACAACGACAACGAACCUGCUCAGACAUCAGACAUUAGACUUCAGACACGGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCG ......((((((((.....((......)).....(((((((((.(((.((......)).)))...)))))))))))..))))))......(((.....))) ( -22.80) >DroSim_CAF1 18042 101 - 1 CGACAACGACAACGACAACGAACCUGCUCAGACAUCAGACAUCAGACUUCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCG ......((((((((....))....((.((((((((..(((((...............)))))....))))))))))..))))))......(((.....))) ( -21.36) >DroEre_CAF1 19414 95 - 1 CGACAACG------ACAACGAAGCUGCUCAGACUUCAGACUUCAGACUGCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAGAAUGCG ......((------((((.((((((....)).))))...(.((((((((..(((.....)))..))..)))))).)..))))))......(((.....))) ( -24.50) >DroYak_CAF1 18782 101 - 1 CGACAACGACAACGACAACGAACCUGCUCAGACAUCAGACAUCAGACUUCAGACGCAGUGUCAACAAUGUCUGACGAAUUGUCGCAAAACCGCAAAAUGCG ......((((((((....))....((.((((((((..(((((..(........)...)))))....))))))))))..))))))......(((.....))) ( -21.00) >consensus CGACAACGACAACGACAACGAACCUGCUCAGACAUCAGACAUCAGACUUCAGACACAGUGUCAACAAUGUCUGACGAAUUGUCGAAAAACCGCAAAAUGCG ((((((.................(((.((........))...))).(.(((((((..((....))..))))))).)..))))))......(((.....))) (-18.78 = -18.66 + -0.12)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:29 2006