
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,258,020 – 13,258,230 |
| Length | 210 |
| Max. P | 0.867341 |

| Location | 13,258,020 – 13,258,122 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.56 |
| Mean single sequence MFE | -20.28 |
| Consensus MFE | -19.44 |
| Energy contribution | -19.40 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.96 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.518204 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13258020 102 - 22224390 UAACACAAGCGCAAGUAAAC------------------AGAGAAAAAUGAUGCUCAUUAACAUUAAAAUUUCGCUUCUUCCACUACAGUGGCCAUUGGAAUAAUGGUCGCCCCGACUCCA ........((....))....------------------.(((.....(((((........)))))......................((((((((((...)))))))))).....))).. ( -17.80) >DroSec_CAF1 3697 102 - 1 UCACACAAGCGCAAGUAAAC------------------AGAGAAAAAUGAUGUUCAUUAACAUUAAAAUUUCACUUCUUCCACUACAGUGGCCAAUGGAAUAAUGGUCGCCCCGACUCCA ........((....))....------------------.(((.....(((((((....)))))))......................(((((((.........))))))).....))).. ( -16.90) >DroSim_CAF1 3727 102 - 1 UAACACAAGCGCAAGUAAAC------------------AGAGAAAAAUGAUGUUCAUUAACAUUAAAAUUUCGCUUCUUCCACUACAGUGGCCAUUGGAAUAAUGGUCGCCCCGACUCCA ........((....))....------------------.(((.....(((((((....)))))))......................((((((((((...)))))))))).....))).. ( -19.80) >DroEre_CAF1 5421 102 - 1 UAACACAAGCGCAAGUAAAC------------------AGAGAAAAAUGGUGUUCAUUAGCAUUAAAAUUUCGCUUCUUCCACUACAGUGGCCAUUGUAAUAAUGGCCGCCCCGACUCGA ........((....))....------------------.(((.....(((((......(((...........))).....)))))..((((((((((...)))))))))).....))).. ( -24.80) >DroYak_CAF1 4161 120 - 1 UAACACAAGCGCAAGUAAACAGAAACGGACACACAAACAGAGAAAAAUGAUGUUCAUUAACAUUAAAAUGUCGCUUCUUCCACUACAGUGGCCAUUGUAAUAAUGGUCGCCCCGACUCCA ........((....)).........(((..........((.((....(((((((....))))))).....)).))............((((((((((...)))))))))).)))...... ( -22.10) >consensus UAACACAAGCGCAAGUAAAC__________________AGAGAAAAAUGAUGUUCAUUAACAUUAAAAUUUCGCUUCUUCCACUACAGUGGCCAUUGGAAUAAUGGUCGCCCCGACUCCA ........((....)).......................(((.....(((((((....)))))))......................((((((((((...)))))))))).....))).. (-19.44 = -19.40 + -0.04)

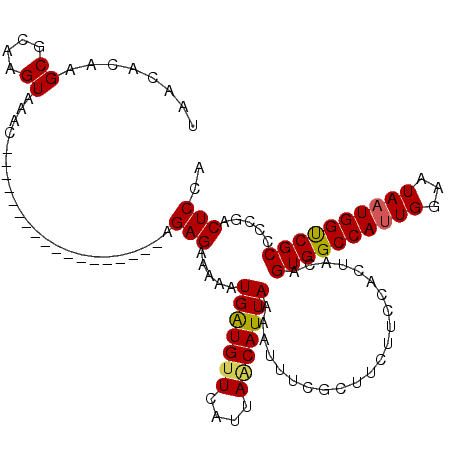

| Location | 13,258,122 – 13,258,230 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.70 |
| Mean single sequence MFE | -23.15 |
| Consensus MFE | -11.60 |
| Energy contribution | -11.56 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.867341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13258122 108 + 22224390 AGUGUCGCAAAUAAAUUCAUUAAAGUGACAUUUUUCAAUUUUAAUGAACAGUACUCGCCAAAAAA-----------AAAAAUGAAGCGCGGGAAAUGGACAUCCAAGA-AAUGAAUACAA .(((((.........(((((((((((((......)).)))))))))))((...((((((......-----------.........).)))))...)))))))......-........... ( -19.16) >DroSec_CAF1 3799 105 + 1 AGUGUCGCAAAUAAAUUCAUUAAAGUGACAUUUUUCUAUUUUAAUGAACAGCACUCACCGAAAAA-----------AAAAA---CGCGCGGGAAAUGGACAUCCAAGA-AAUGAAUACAA .(((((((.......(((((((((((((......)).)))))))))))..((.............-----------.....---.))))).....(((....)))...-.....)))).. ( -16.81) >DroSim_CAF1 3829 104 + 1 AGUGUCGCAAAUAAAUUCAUUAAAGUGACAUUUUUCAAUUUUAAUGAACAGCACUCGCCGAAAA------------AAAAA---CGCGCGGGAAAUGGACAUCCAAGA-AAUGAAUACAA .(((((.........(((((((((((((......)).)))))))))))((...(((((((....------------.....---)).)))))...)))))))......-........... ( -21.70) >DroEre_CAF1 5523 115 + 1 AGUGUCGCAAAUAAAUUCAUUAAAGUGGCAUUUUUCAAUUUUAAUGAACACCAUCC---GACGAGCG-AGCUGCCCAAAAAGCUCGCUCCCCAAAUGGACUUCCAAGA-AAUGAAUACAA ...((((........(((((((((((...........))))))))))).......)---)))(((((-((((........)))))))))......(((....)))...-........... ( -31.56) >DroYak_CAF1 4281 116 + 1 AGUGUCGCAAAUAAAUUCAUUAAAGUGGCAUUUUUCAAUUUUAAUGAACAGCCCCCACCCAAAAGCGCAAAAGCGCAAAAA-CGCGG---GGAAAUGGACUUGCAAGAAAAUGAAUACAA ......((((.....(((((((((((...........)))))))))))((..((((.(......((((....)))).....-.).))---))...))...))))................ ( -26.50) >consensus AGUGUCGCAAAUAAAUUCAUUAAAGUGACAUUUUUCAAUUUUAAUGAACAGCACUCACCGAAAAA___________AAAAA___CGCGCGGGAAAUGGACAUCCAAGA_AAUGAAUACAA .((((((........(((((((((((((......)).)))))))))))...............................................(((....)))......)).)))).. (-11.60 = -11.56 + -0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:14 2006