
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,971,710 – 12,971,844 |
| Length | 134 |
| Max. P | 0.848604 |

| Location | 12,971,710 – 12,971,808 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.03 |
| Mean single sequence MFE | -24.02 |
| Consensus MFE | -14.64 |
| Energy contribution | -15.45 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.696068 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
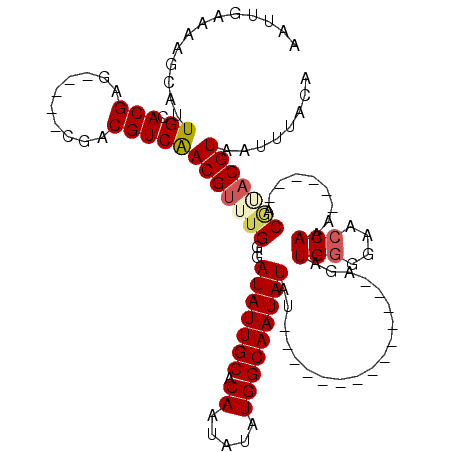
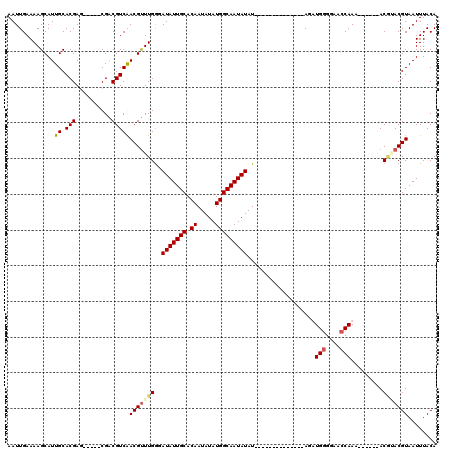
>X_DroMel_CAF1 12971710 98 + 22224390 AAUUGAAAAGCAUUGCACGAG-----CGUCGUCGACGCUUGGGAUAUUGCACAAUAUAUGGCAAUAUAU--------------AGAUGUGGAACCAAA---CAGACGUACGUAAUUUACA ...........(((((.((((-----((((...))))))))((...((.((((.((((((....)))))--------------)..)))).))))...---.........)))))..... ( -22.30) >DroSim_CAF1 6713 101 + 1 AAUUGAAAAGCAUUGCACGAG-----CGACGUCGACGUUUGGGAUAUUGCACAAUAUAUGGCAAUAUAU--------------AGAUGGGGAACCAAACAACAGACGUACGUAAUUUACA .........((.((((....)-----))).))(((((((((..(((((((.((.....)))))))))..--------------...(((....))).....))))))).))......... ( -24.10) >DroEre_CAF1 5949 94 + 1 AAUUGAAAAGCAUUGCACGAG-----CGACGUCAACGUUUGGUAUAUUGCACAAUAUAUGGCAAUAUAU--------------AGUUGGGGAACCAA-------ACAAACGUAAUUUACA .........((.((((....)-----))).))..((((((((((((((((.((.....)))))))))))--------------..((((....))))-------.)))))))........ ( -26.30) >DroYak_CAF1 6289 114 + 1 AAUUGAAAAGCAUUGCACGAACGGAACGACGUCAACGUUUGAGAUAUUGCACAAUAUAUGGCAAUAUAUCUAUGUAUAUGUACAGUUGGGGAACCAAA------ACCGACGUAAUUUACA .......................(((..(((((.((((..((.(((((((.((.....))))))))).)).))))..........((((....)))).------...)))))..)))... ( -23.40) >consensus AAUUGAAAAGCAUUGCACGAG_____CGACGUCAACGUUUGGGAUAUUGCACAAUAUAUGGCAAUAUAU______________AGAUGGGGAACCAAA______ACGUACGUAAUUUACA .............((.(((..........)))))(((((((..(((((((.((.....)))))))))...................(((....))).........)))))))........ (-14.64 = -15.45 + 0.81)

| Location | 12,971,745 – 12,971,844 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 80.62 |
| Mean single sequence MFE | -24.30 |
| Consensus MFE | -14.88 |
| Energy contribution | -15.50 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.848604 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
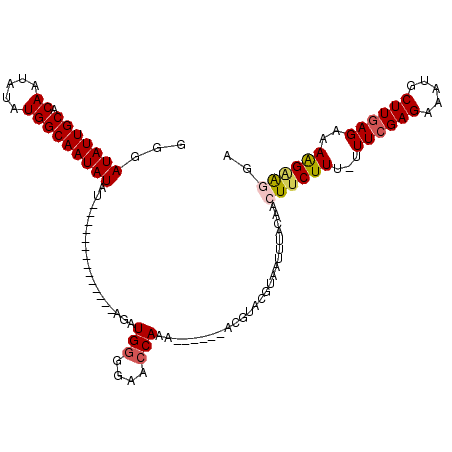
>X_DroMel_CAF1 12971745 99 + 22224390 GGGAUAUUGCACAAUAUAUGGCAAUAUAU--------------AGAUGUGGAACCAAA---CAGACGUACGUAAUUUACAACUUCUUU-UUUCGAGAAAUGCUUGAGAAAGGAGUGA .((...((.((((.((((((....)))))--------------)..)))).))))...---...................(((.((((-(((((((.....)))))))))))))).. ( -20.90) >DroSim_CAF1 6748 102 + 1 GGGAUAUUGCACAAUAUAUGGCAAUAUAU--------------AGAUGGGGAACCAAACAACAGACGUACGUAAUUUACAACUUCUUU-UUUCGAGAAAUGCUUGAGAAAGGAGCGA ...(((((((.((.....)))))))))..--------------...(((....))).........(((..(((...)))....(((((-(((((((.....))))))))))))))). ( -25.00) >DroEre_CAF1 5984 95 + 1 GGUAUAUUGCACAAUAUAUGGCAAUAUAU--------------AGUUGGGGAACCAA-------ACAAACGUAAUUUACAACUUCUUU-UUUCGAGAAAUGCUUGAGUUAAGUAGGU .(((((((((.((.....)))))))))))--------------..((((....))))-------................(((((((.-.((((((.....))))))..))).)))) ( -23.60) >DroYak_CAF1 6329 111 + 1 GAGAUAUUGCACAAUAUAUGGCAAUAUAUCUAUGUAUAUGUACAGUUGGGGAACCAAA------ACCGACGUAAUUUACAACUUCUUUUUUUCGAGAAAUGCUUAAGUUAAGGAGGU ((.(((((((.((.....))))))))).))..(((((((((....((((....)))).------....)))))...))))((((((((..((..((.....))..))..)))))))) ( -27.70) >consensus GGGAUAUUGCACAAUAUAUGGCAAUAUAU______________AGAUGGGGAACCAAA______ACGUACGUAAUUUACAACUUCUUU_UUUCGAGAAAUGCUUGAGAAAAGAAGGA ...(((((((.((.....)))))))))...................(((....))).........................((((((...((((((.....))))))..)))))).. (-14.88 = -15.50 + 0.62)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:37:17 2006