
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,948,791 – 12,948,904 |
| Length | 113 |
| Max. P | 0.814892 |

| Location | 12,948,791 – 12,948,904 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 92.77 |
| Mean single sequence MFE | -40.03 |
| Consensus MFE | -33.94 |
| Energy contribution | -35.62 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.814892 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12948791 113 + 22224390 AGGGUAUCAACAGGUGCUCAAUUUCCACCUGUGCGCACUGCGCUGUUGUGAUUUUGUUGCCGCGACGGCGCAGCUUUUGCUUUUGACACAUUUUCGCGGCUGUCUCGCCGCCG ...((.((((((((((.........)))))))..((((((((((((((((..........)))))))))))))....)))...))).))......(((((......))))).. ( -44.60) >DroSec_CAF1 7248 107 + 1 AGGGUAUCAACAGGUGCUCAAUUUCCACCUGUGCGCACAGCGCUGUUGUGAUUUUGUUGCCGCGACGGCGCAGCUUUUGCUUUUGACACAUUU------CUGUCUCGCCGCCG ..(((....(((((((.........)))))))(((....(((((((((((..........)))))))))))(((....)))...((((.....------.)))).))).))). ( -37.20) >DroSim_CAF1 8005 113 + 1 AGGGUAUCAACAGGUGCUAAAUUUCCACCGGUGCGCACAGCGCUGUUGUGAUUUUGUUACCGCGACGGCGCAGCUUUUGCUUUUGACACAUUUUCGCGGCUGUCUCGCCGCCG .(.(((((....((((.........))))))))).)...(((((((((((..........)))))))))))(((....)))..............(((((......))))).. ( -38.10) >DroEre_CAF1 6488 112 + 1 AGGGUAUCAACAGGUGCUCAAUUUCCACCUGUGCCCUCUGCGUUGUUGUGA-UUUGCUGCCGCGACGGCGCAGCUUUUGCUUUUGACACAUUUUCGCGGCUGUCUCGUCGCCG (((((....(((((((.........))))))))))))(((((((((((((.-........)))))))))))))......................(((((......))))).. ( -40.20) >consensus AGGGUAUCAACAGGUGCUCAAUUUCCACCUGUGCGCACAGCGCUGUUGUGAUUUUGUUGCCGCGACGGCGCAGCUUUUGCUUUUGACACAUUUUCGCGGCUGUCUCGCCGCCG ...((.((((((((((.........)))))))(((..(((((((((((((..........)))))))))))))....)))...))).))......(((((......))))).. (-33.94 = -35.62 + 1.69)

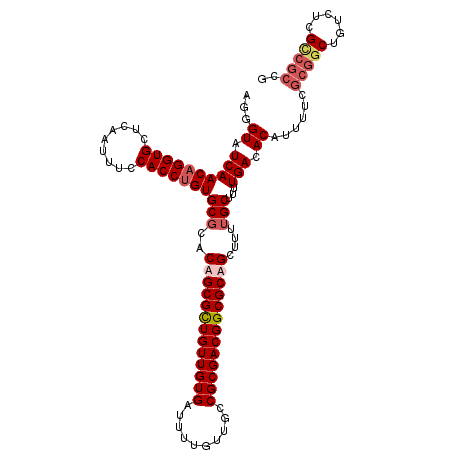
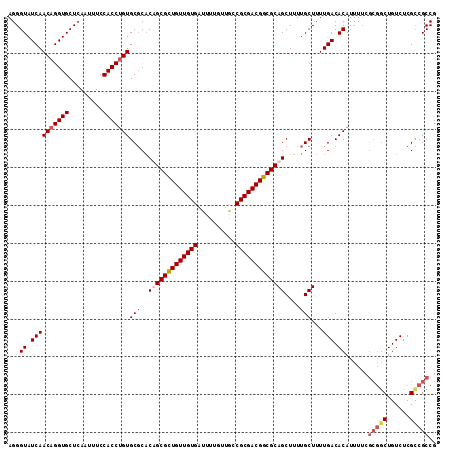
| Location | 12,948,791 – 12,948,904 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 92.77 |
| Mean single sequence MFE | -37.71 |
| Consensus MFE | -29.00 |
| Energy contribution | -31.50 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.668732 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12948791 113 - 22224390 CGGCGGCGAGACAGCCGCGAAAAUGUGUCAAAAGCAAAAGCUGCGCCGUCGCGGCAACAAAAUCACAACAGCGCAGUGCGCACAGGUGGAAAUUGAGCACCUGUUGAUACCCU ..(((((......)))))......((((((...((....(((((((.((.(.((........)).).)).)))))))..))(((((((.........)))))))))))))... ( -41.40) >DroSec_CAF1 7248 107 - 1 CGGCGGCGAGACAG------AAAUGUGUCAAAAGCAAAAGCUGCGCCGUCGCGGCAACAAAAUCACAACAGCGCUGUGCGCACAGGUGGAAAUUGAGCACCUGUUGAUACCCU (((((((((.(((.------...))).))....(((.....)))))))))).(....)...((((.....((((...))))(((((((.........)))))))))))..... ( -39.50) >DroSim_CAF1 8005 113 - 1 CGGCGGCGAGACAGCCGCGAAAAUGUGUCAAAAGCAAAAGCUGCGCCGUCGCGGUAACAAAAUCACAACAGCGCUGUGCGCACCGGUGGAAAUUUAGCACCUGUUGAUACCCU ..(((((......)))))......(((((((.(((....)))((((((.(((.((............)).))).)).))))...((((.........))))..)))))))... ( -35.40) >DroEre_CAF1 6488 112 - 1 CGGCGACGAGACAGCCGCGAAAAUGUGUCAAAAGCAAAAGCUGCGCCGUCGCGGCAGCAAA-UCACAACAACGCAGAGGGCACAGGUGGAAAUUGAGCACCUGUUGAUACCCU ((....)).....(((((((....((((....(((....)))))))..))))))).((...-..........))..((((.(((((((.........))))))).....)))) ( -34.52) >consensus CGGCGGCGAGACAGCCGCGAAAAUGUGUCAAAAGCAAAAGCUGCGCCGUCGCGGCAACAAAAUCACAACAGCGCAGUGCGCACAGGUGGAAAUUGAGCACCUGUUGAUACCCU ..(((((......)))))......((((((...((....(((((((.((.(.((........)).).)).)))))))..))(((((((.........)))))))))))))... (-29.00 = -31.50 + 2.50)

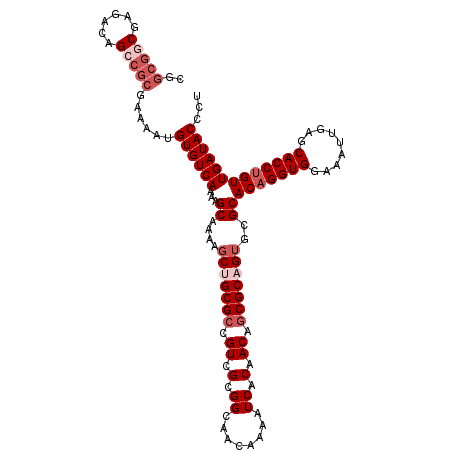
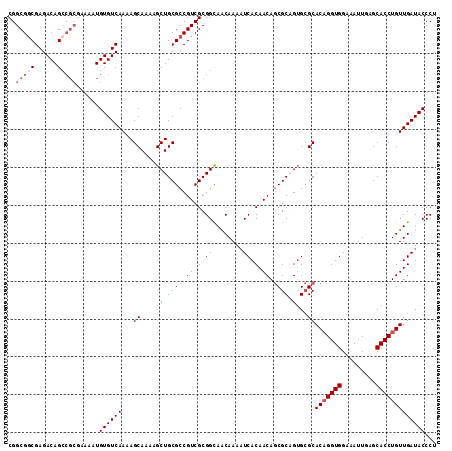
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:36:52 2006