| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,417,163 – 1,417,263 |
| Length | 100 |
| Max. P | 0.881617 |

| Location | 1,417,163 – 1,417,263 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 81.69 |
| Mean single sequence MFE | -30.84 |
| Consensus MFE | -18.12 |
| Energy contribution | -18.60 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.630469 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
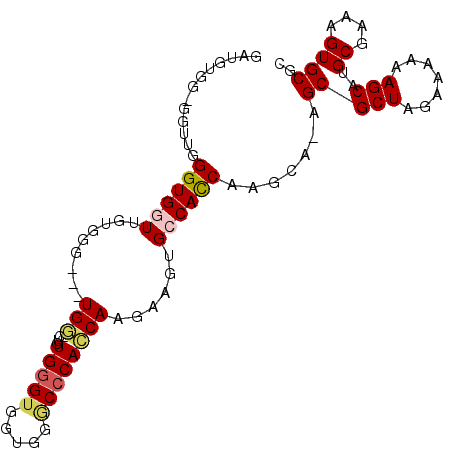
>X_DroMel_CAF1 1417163 100 + 22224390 GAUGUGGAGGUUGGGUGGUUGUGGGUGGUGGGUGGUGGGUGGUGGGCCCACCAAGAAGUGCCACCAAGCA-AGCGCUAGAAAAAAGCAUGCGAAAGUGCGC .((((.....((.((((.((((.((((((...((((((((.....))))))))......))))))..)))-).)))).)).....))))((....)).... ( -39.40) >DroSec_CAF1 23631 94 + 1 GAUGUGGGGGUUGGGUGAUUG--GG---UGG-CUGUGAGUGGUGGGCCCACCAAGAAGUGCCACCAGGCA-AGCGCUAGAAAAAAGCAUGCGAAAGUGCGC .((((.....((.((((.(((--((---(((-(......(((((....)))))......))))))...))-).)))).)).....))))((....)).... ( -31.10) >DroSim_CAF1 14252 95 + 1 GAUGUGGGGGUUGGGUGAUUG--GG---UGG-CUGUGGGUGGUGGGCCCACCAAGAAGUGCCACCAAGCAAAGCGCUAGAAAAAAGCAUGCGAAAGUGCGC ......((.(((..((.....--((---(((-(......(((((....)))))......))))))..))..))).))........((..((....))..)) ( -32.40) >DroEre_CAF1 18940 89 + 1 G-CGU-----AUGGGUGGUCGUGGG---UGU-UCGUGGGUGGUGGACCCAGCAAGAAGUGCCACC-AGCA-AGCGCUAGAAAAAAGCAUGCGAAAGUGCGC (-(((-----((.((((((......---(((-...(((((.....))))))))......))))))-.(((-.((...........)).)))....)))))) ( -30.40) >DroYak_CAF1 15164 83 + 1 -------------GGUGGUGAUGGG---UGG-AAUUGGGCGGUGGACCCAGCAAGAAGUGCCAUCAAGCA-AGCGCUAGAAAAAAGCAUGCGAAAGUGCGC -------------.((..((((((.---...-...(((((....).))))((.....)).)))))).)).-.(((((.......)))..((....)))).. ( -20.90) >consensus GAUGUGG_GGUUGGGUGGUUGUGGG___UGG_CUGUGGGUGGUGGGCCCACCAAGAAGUGCCACCAAGCA_AGCGCUAGAAAAAAGCAUGCGAAAGUGCGC .............((((((.........(((....(((((.....))))))))......)))))).......(((((.......)))..((....)))).. (-18.12 = -18.60 + 0.48)

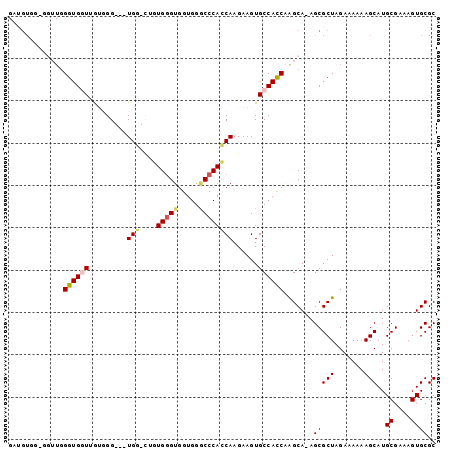
| Location | 1,417,163 – 1,417,263 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 81.69 |
| Mean single sequence MFE | -22.28 |
| Consensus MFE | -16.38 |
| Energy contribution | -16.66 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.881617 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
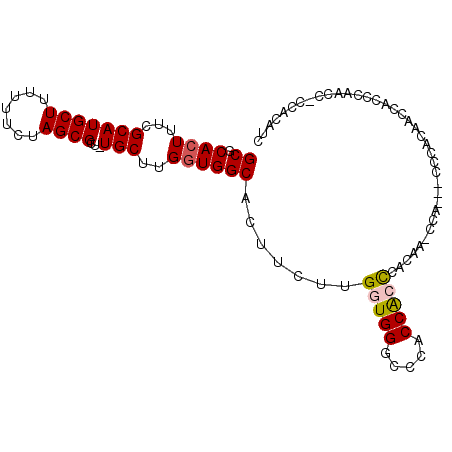
>X_DroMel_CAF1 1417163 100 - 22224390 GCGCACUUUCGCAUGCUUUUUUCUAGCGCU-UGCUUGGUGGCACUUCUUGGUGGGCCCACCACCCACCACCCACCACCCACAACCACCCAACCUCCACAUC ((((......))..(((.......))))).-((..((((((.......(((((((.......))))))).))))))..))..................... ( -24.80) >DroSec_CAF1 23631 94 - 1 GCGCACUUUCGCAUGCUUUUUUCUAGCGCU-UGCCUGGUGGCACUUCUUGGUGGGCCCACCACUCACAG-CCA---CC--CAAUCACCCAACCCCCACAUC ((((......))..(((.......))))).-.....((((((......(((((....)))))......)-)))---))--..................... ( -24.30) >DroSim_CAF1 14252 95 - 1 GCGCACUUUCGCAUGCUUUUUUCUAGCGCUUUGCUUGGUGGCACUUCUUGGUGGGCCCACCACCCACAG-CCA---CC--CAAUCACCCAACCCCCACAUC ((((......))..(((.......))))).......((((((......(((((....)))))......)-)))---))--..................... ( -24.30) >DroEre_CAF1 18940 89 - 1 GCGCACUUUCGCAUGCUUUUUUCUAGCGCU-UGCU-GGUGGCACUUCUUGCUGGGUCCACCACCCACGA-ACA---CCCACGACCACCCAU-----ACG-C (((.......(((((((.......))))..-))).-(((((..........(((((.....)))))((.-...---....)).)))))...-----.))-) ( -20.10) >DroYak_CAF1 15164 83 - 1 GCGCACUUUCGCAUGCUUUUUUCUAGCGCU-UGCUUGAUGGCACUUCUUGCUGGGUCCACCGCCCAAUU-CCA---CCCAUCACCACC------------- ((((......))..(((.......))))).-....((((((((.....)))(((((.....)))))...-...---..))))).....------------- ( -17.90) >consensus GCGCACUUUCGCAUGCUUUUUUCUAGCGCU_UGCUUGGUGGCACUUCUUGGUGGGCCCACCACCCACAA_CCA___CCCACAACCACCCAACC_CCACAUC ((.((((...(((((((.......))))...)))..)))))).......(((((.....)))))..................................... (-16.38 = -16.66 + 0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:45:49 2006